Fragment Merging – and Flipping – on the Leucine Zipper of MITF
•January 26, 2026
0
Why It Matters
Demonstrating ligandability of a leucine‑zipper transcription factor opens new avenues for drugging previously ‘undruggable’ cancer targets and informs emerging PROTAC strategies.
Key Takeaways
- •19F NMR screened MITF DBD against LEF4000 library
- •Only nine hits; hit rate among lowest observed
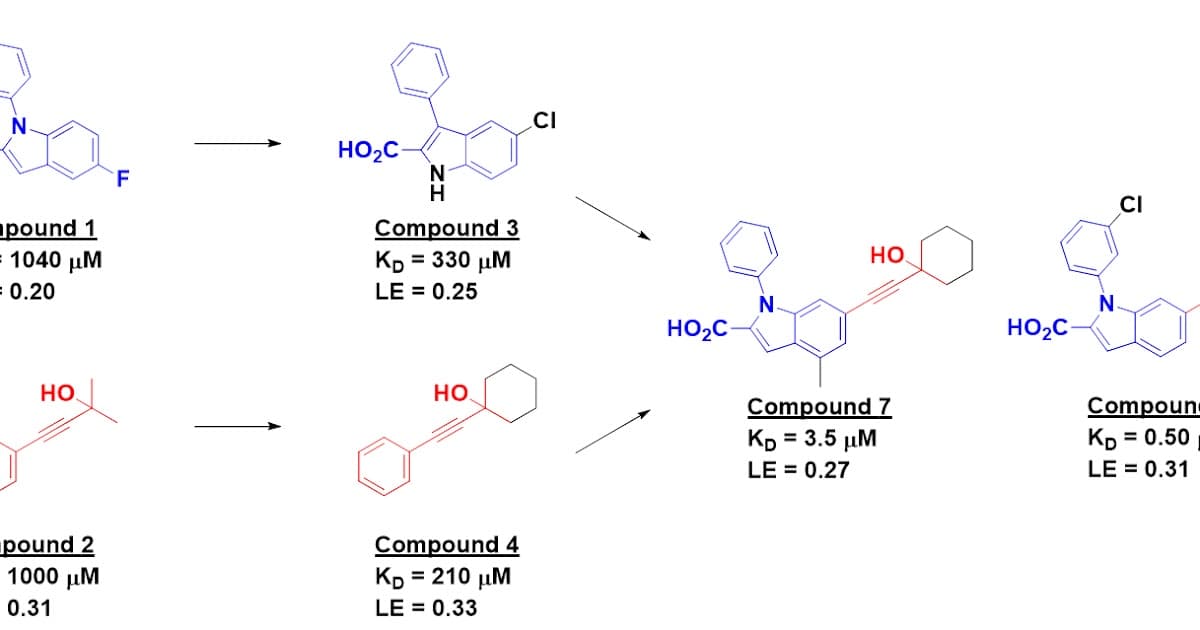
- •Fragment merging produced 100× tighter binder
- •Compounds 7 and 8 achieve sub‑micromolar affinity
- •Binding does not yet affect DNA interaction or cells
Pulse Analysis
Transcription factors with leucine‑zipper motifs have long been considered intractable because their coiled‑coil surfaces lack deep pockets for small‑molecule engagement. MITF, a master regulator of melanocyte development and melanoma progression, exemplifies this challenge. By focusing on the DNA‑binding domain’s kinked helix, the Novartis team leveraged 19F‑based fragment screening—a technique that excels at detecting weak binders in high‑throughput settings—despite the target’s predominantly disordered nature. The effort underscores how modern biophysical tools can illuminate even the most elusive protein interfaces.
The screen produced a modest nine‑hit set, yet two chemical series were advanced through iterative analog testing. Crystallography confirmed that fragments 3 and 4 occupied a previously uncharacterized kink pocket, enabling a rational merge that amplified binding affinity by two orders of magnitude. Subsequent optimization delivered compounds 7 and 8 with low‑nanomolar to sub‑micromolar dissociation constants measured by ITC. Notably, compound 8 adopts a 180° flipped orientation relative to 7, a phenomenon documented in other fragment‑based campaigns and indicative of the plasticity inherent in shallow protein pockets.
While the ligands do not yet perturb MITF’s DNA‑binding activity or demonstrate cellular efficacy, the study validates the concept that leucine‑zipper dimers can be drugged. Future work may explore covalent warheads to boost potency or repurpose these fragments as recruitment elements in PROTACs and molecular glues. As the field seeks to expand the druggable proteome, the MITF case provides a template for tackling similarly feature‑poor transcription factors, suggesting that strategic fragment merging and structural insight can overcome longstanding barriers.
Fragment merging – and flipping – on the leucine zipper of MITF
0
Comments
Want to join the conversation?
Loading comments...