
Two Plugins Transform Claude Code Web Development
1/7 Most people use Claude Code wrong for building websites. They type "build me a landing page" and get a generic Bootstrap mess. Then they spend hours fixing it. I did this too until I started using two plugins that completely changed my workflow.

Never Trust Bulk RNA‑seq without These 7 QC Checks
1/9 Every bulk RNA-seq experiment I run goes through the same 7 checks before I trust the results. I've been burned enough times to know: if you skip QC, you will find out the hard way. Usually during a meeting with...

Biologists Wielding AI Will Outpace Those Who Don’t
1/ AI won't replace you. But a biologist using AI will. Especially in bioinformatics, where the questions never stop coming. https://t.co/ckfZWjnei3

Claude’s /Btw Lets You Ask Quick Questions Discreetly
1/ TIL about /btw in Claude Code and I wish I had this weeks ago. You can ask Claude a quick question while it's still working on something else. The answer doesn't pollute your conversation history. https://t.co/FmPNX0EX6Y

Top Resources for TF Binding, Enhancers, Histone Marks
8 Resources to study Transcription factor binding, enhancers and histone modification distribution 1. ENCODE https://t.co/N5hScyoAoP

Anthropic's 33‑page Claude Skill Guide: Key Takeaways
Anthropic just dropped a 33-page guide on building skills for Claude. I read the whole thing. Here's what actually matters: https://t.co/GbYYiAYp9b

Claude Code Runs Playwright, Revolutionizing Web App Testing
Claude Code can't see your browser. But it can run Playwright. And that changes everything for testing web apps. https://t.co/PzWx3D8ew6 https://t.co/dDlQLV2S7P

Three Content Buckets Are Enough for Daily Posts
Someone asked me at a career workshop: "How do you come up with stuff to post every day?" My answer: you only need three buckets. That's it.

Open-Source Claude Code Setup: Full Toolkit Revealed
Boris Cherny created Claude Code. Then he open-sourced his entire setup for using it. Memory files, subagents, skills, the whole thing. Here's what's inside and why you should steal it:

DNA Repair Mutations Predict Response to Chemo‑Immunotherapy in Pancreatic Cancer
DNA Repair gene alterations and efficacy from gemcitabine and nab-paclitaxel with/without durvalumab and tremelimumab in metastatic pancreatic ductal adenocarcinoma https://t.co/v39bOreJwu

Coding Beats Pipettes: The Biologist’s Ultimate Lab Tool
1/ Your most powerful lab tool isn't a pipette. It's a keyboard. If you're a biologist who hasn't learned to code yet, here are 10 rules that changed everything for me: https://t.co/DhSXBXGitk

Stop Claude Code’s 3‑second Permission Prompts—Quick Fixes
Claude Code keeps asking for permission every 3 seconds and you're about to lose your mind. Here's how to fix it — from nuclear to surgical:

DBiTplus Merges Imaging and Sequencing Spatial Omics
Integration of imaging-based and sequencing-based spatial omics mapping on the same tissue section via DBiTplus https://t.co/YtsDFQNt9H https://t.co/ffKPWLDViY
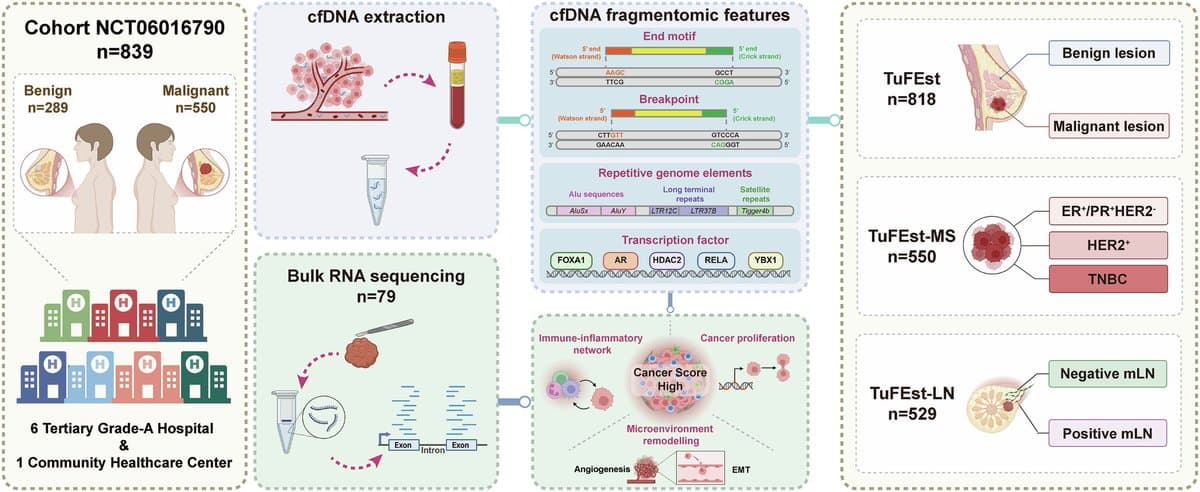
Fragmentomic Liquid Biopsy Detects Early Breast Cancer, Subtypes, Nodes
Fragmentomic liquid biopsy enables early breast cancer detection, molecular subtyping and lymph node assessment https://t.co/RC5rTrFO4l https://t.co/s0Z6lc7Jeb

AlphaFold Isn’t the Drug‑development Silver Bullet
Even with AlphaFold, protein structure is not a solved problem. And protein structure was never the bottleneck for drug development anyway. Let me explain.
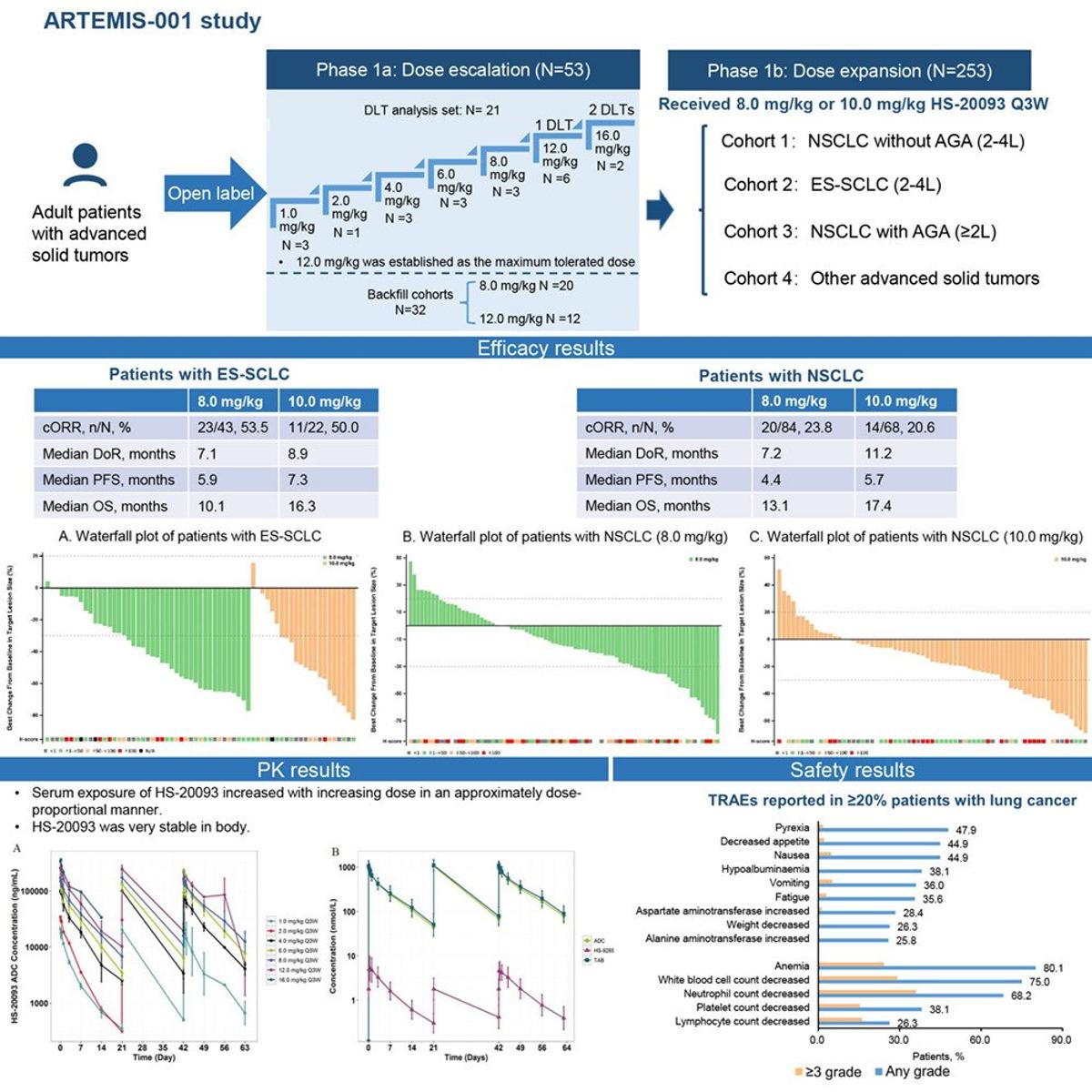
B7‑H3 ADC HS‑20093 Shows Early Lung Cancer Activity
HS-20093, a B7-H3-targeted antibody-drug conjugate in lung cancer: Results from the ARTEMIS-001 phase 1a/b trial https://t.co/Jr90iovVeo https://t.co/Nl2WkprjU3
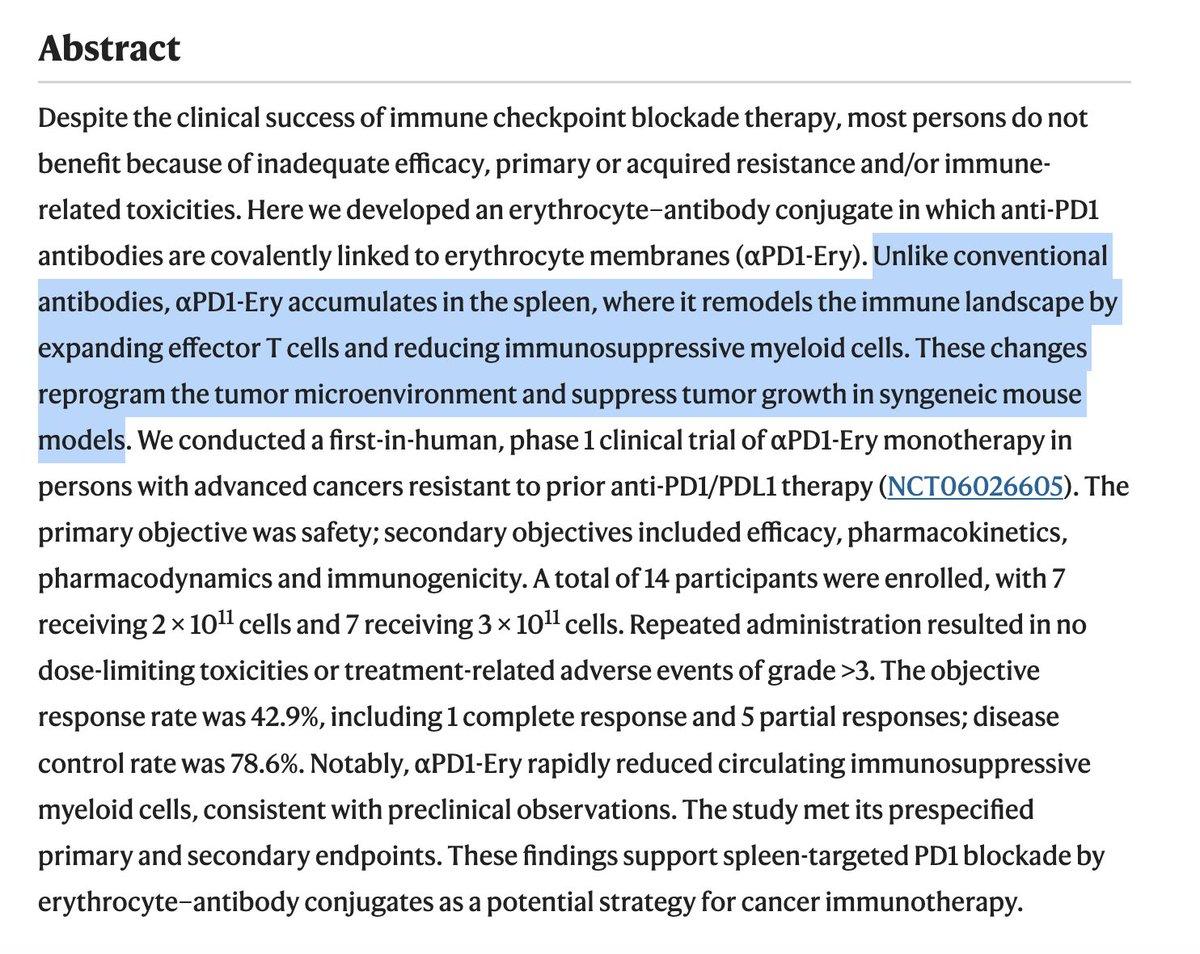

Red Blood Cell–PD1 Conjugates Show Promise in Resistant Tumors
Erythrocyte–anti-PD1 conjugates in persons with advanced solid tumors resistant to anti-PD1/PDL1: preclinical characterization and results of a phase 1 trial https://t.co/1GQMlUHn4Y https://t.co/0it5nTvUm0

Sampling Timing Is as Critical as Measurement
When you drew the sample matters as much as what you measured. Biological data has nuances that get buried if you don't think about timing. https://t.co/eWzEu8FDfc
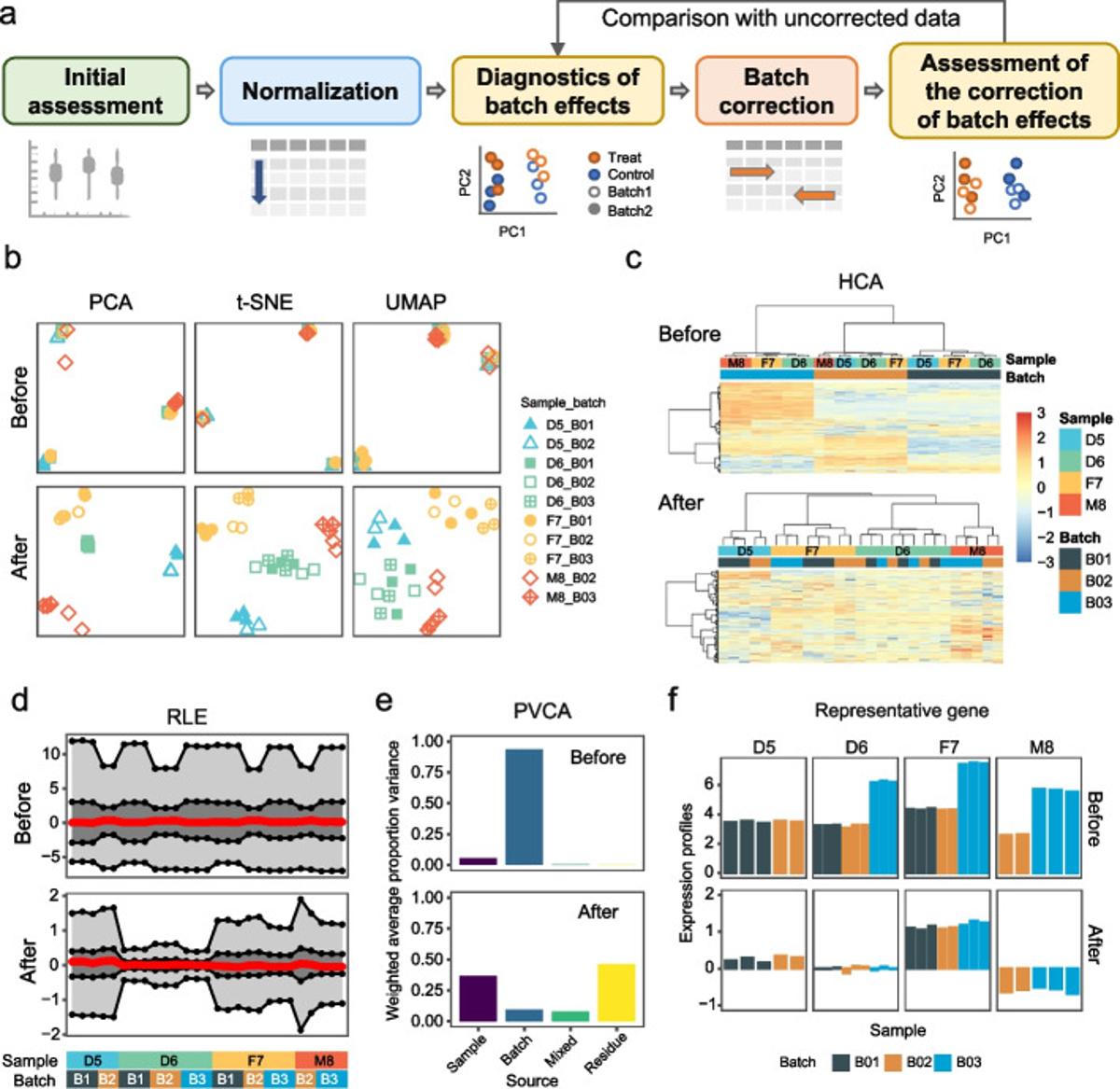
Batch Effects Misclassified 162 Patients, Causing Unnecessary Chemo
Batch effects once caused 162 patients to be misclassified. 28 of them received incorrect or unnecessary chemotherapy. The culprit? Contaminated RNA extraction that introduced technical artifacts into the data. https://t.co/WBBFKgvzVC

Benchmarking Single-Cell Annotation Overlooks Real-World Scale
Why I think your single-cell cell annotation benchmarking is missing the mark 👇 You trained your model on large of number of cells (millions), and you use your model to annotate a new dataset. https://t.co/GHpQkfLtOE

Integrating Omics Data, Not Algorithms, Is Pharma’s Biggest Challenge
The hardest problem in pharma data science isn't the algorithm. It's getting three different omics datasets to agree with each other. That's exactly what we'll dig into at #ddpeast26 on June 10 in Waltham. I'm joining as a panelist and roundtable moderator....

Biologists Using AI Will Outpace Those Who Don't
1/ AI won't replace you. But a biologist who uses AI will. Especially in bioinformatics, where the questions never stop coming. https://t.co/q2OHzntcoV

Large cfDNA Methylome Atlas Powers Multi‑Cancer Detection
A pan-cancer compendium of 1,294 plasma cell-free DNA methylomes and fragmentomes enabling multicancer detection https://t.co/A6gHwcEzqq https://t.co/mSMcR0a833

Single-Cell Atlas Links Marrow Immune Dysregulation to Myeloma Outcomes
A single-cell atlas characterizes dysregulation of the bone marrow immune microenvironment associated with outcomes in multiple myeloma https://t.co/5t0M2eX0fB https://t.co/p3kv8LEzrC

Bioinformatics' Toughest Skill: Bridging the Biologist Gap
1/ The hardest skill in bioinformatics? It's not coding. It's not stats. A hiring manager told me their biggest challenge: finding bioinformaticians who can talk to biologists. Here's why that matters: https://t.co/76HjMDdGIK
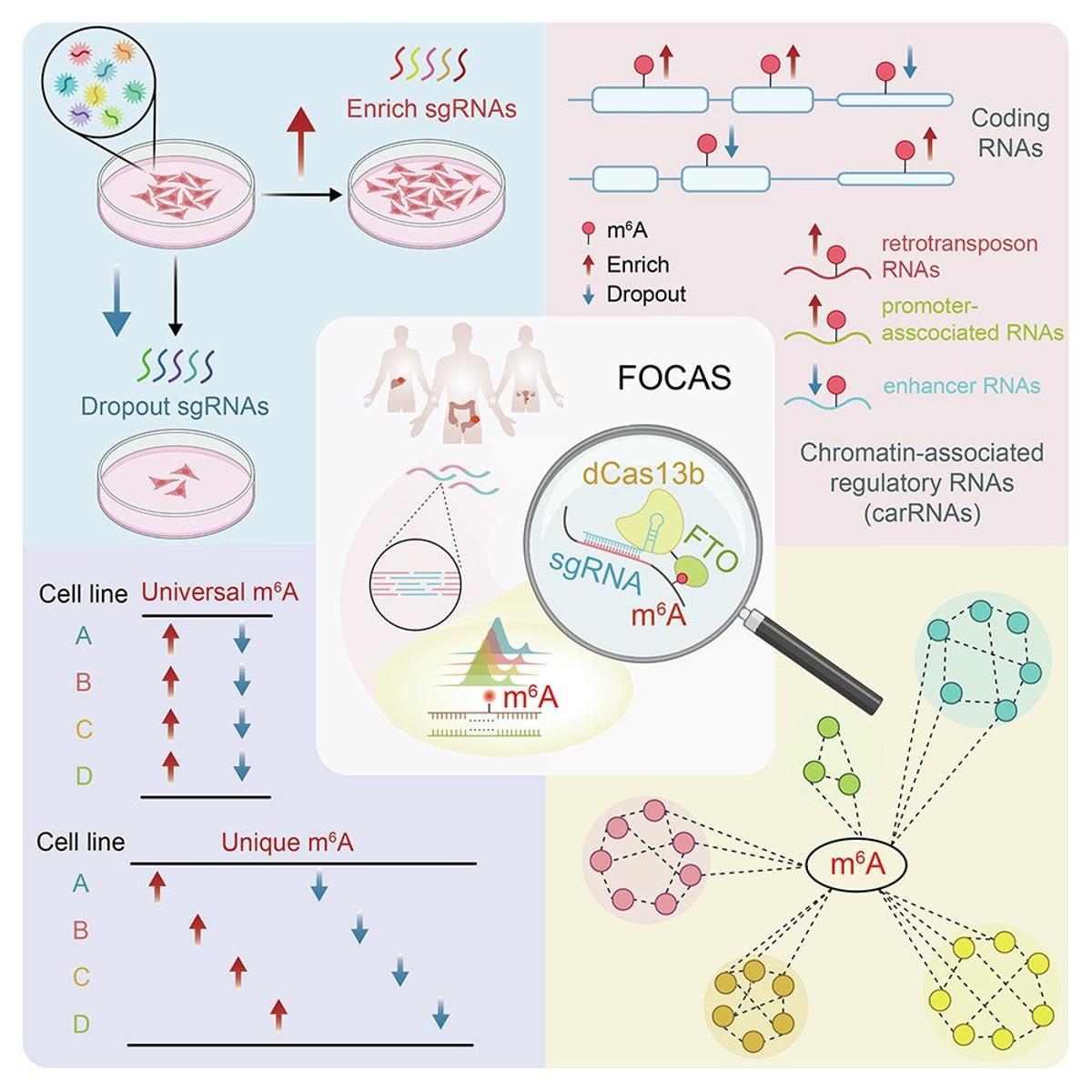
FOCAS Enables Precise Site‑Specific M6A Erasure via CRISPR
m6A is the most common internal mRNA modification, but we still don't know what most m6A sites actually DO. Zhang et al. built FOCAS -- a CRISPR-based platform using dCas13b-FTO to precisely remove m6A at specific sites without touching the DNA. https://t.co/G6kSgpSSbV...

Learn 6 Plot Types to Replicate Any Genomics Figure
Master these 6 types of plots to reproduce any genomics paper figures. I will prove it at the end of the video. https://t.co/gjRs0gY6Mx

Bioinformatics Needs Git: 6 Essential Commands
1/ If you're doing bioinformatics without Git, you're gambling with your research. Here are 6 Git commands every bioinformatician must know 🧵 https://t.co/gFhsTYIsMv

Single-Cell RNA‑seq Isn’t Always Necessary—Choose Wisely
1/ 🧵 No, you do not always Need Single-Cell RNA-seq Single-cell RNA-seq is popular and powerful, but it's not always the best choice. Here's why you should think carefully before diving in. https://t.co/E9DdeHI4b6

Detected Gene Count Isn’t a True Activity Metric
1/ Early scRNA‑seq papers sometimes treated “number of detected genes per cell” as a direct biological readout: more genes = more active cell, fewer genes = quiescent or distinct state. https://t.co/IjLWrR4uu7
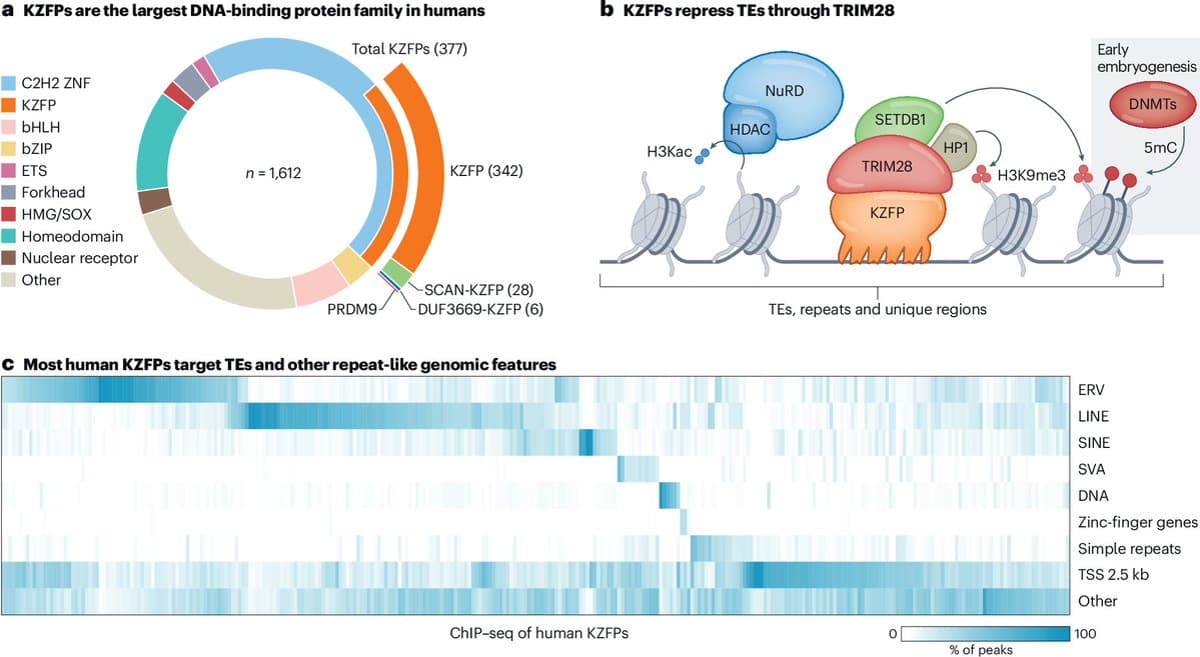
KRAB Zinc‑Finger Proteins Boost Transposable Element Domestication
The role of KRAB zinc-finger proteins in expanding the domestication potential of transposable elements https://t.co/4VtCDQomg6

Sex Chromosomes Cause Mapping Artifacts in Whole‑genome Sequencing
1/ Sex chromosomes are bias magnets in WGS. Early large-scale genomes often showed weird copy-number or coverage patterns on X and Y that looked like biology, but were really mapping artifacts. https://t.co/YnApmtxHqm
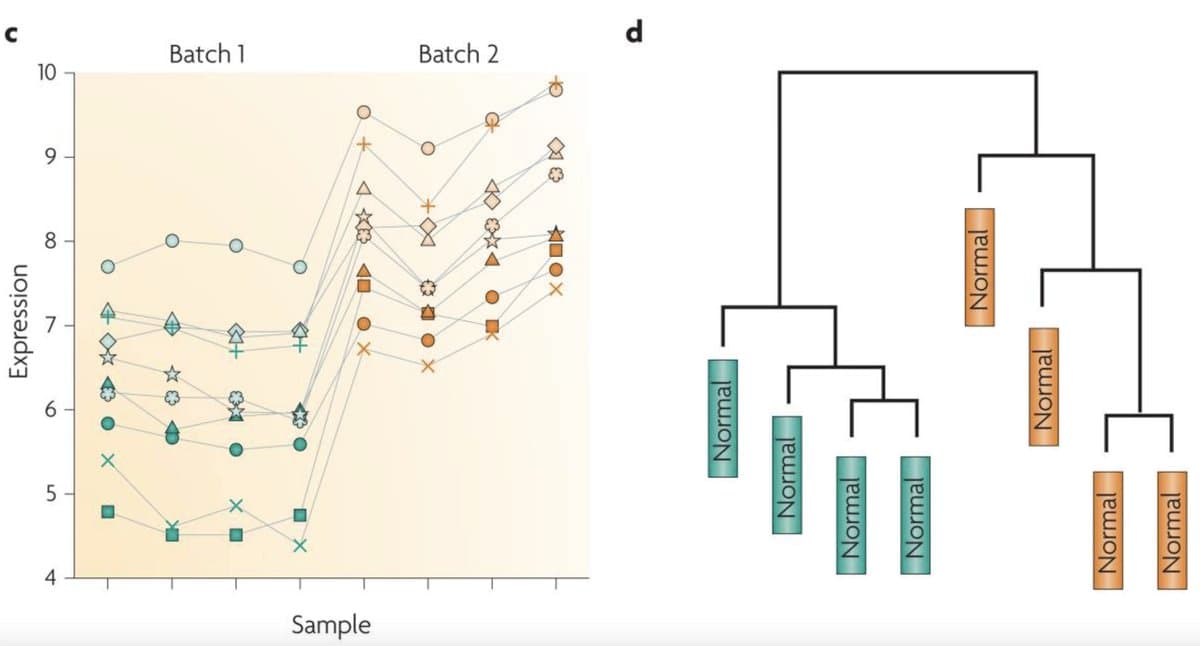
RNA‑seq Batch Effects Can Fabricate False Biological Results
1/ RNA‑seq batch effects are one of the easiest ways to fool yourself in genomics. They can create beautiful, completely wrong biology if you’re not careful. https://t.co/VBtySwZHVL
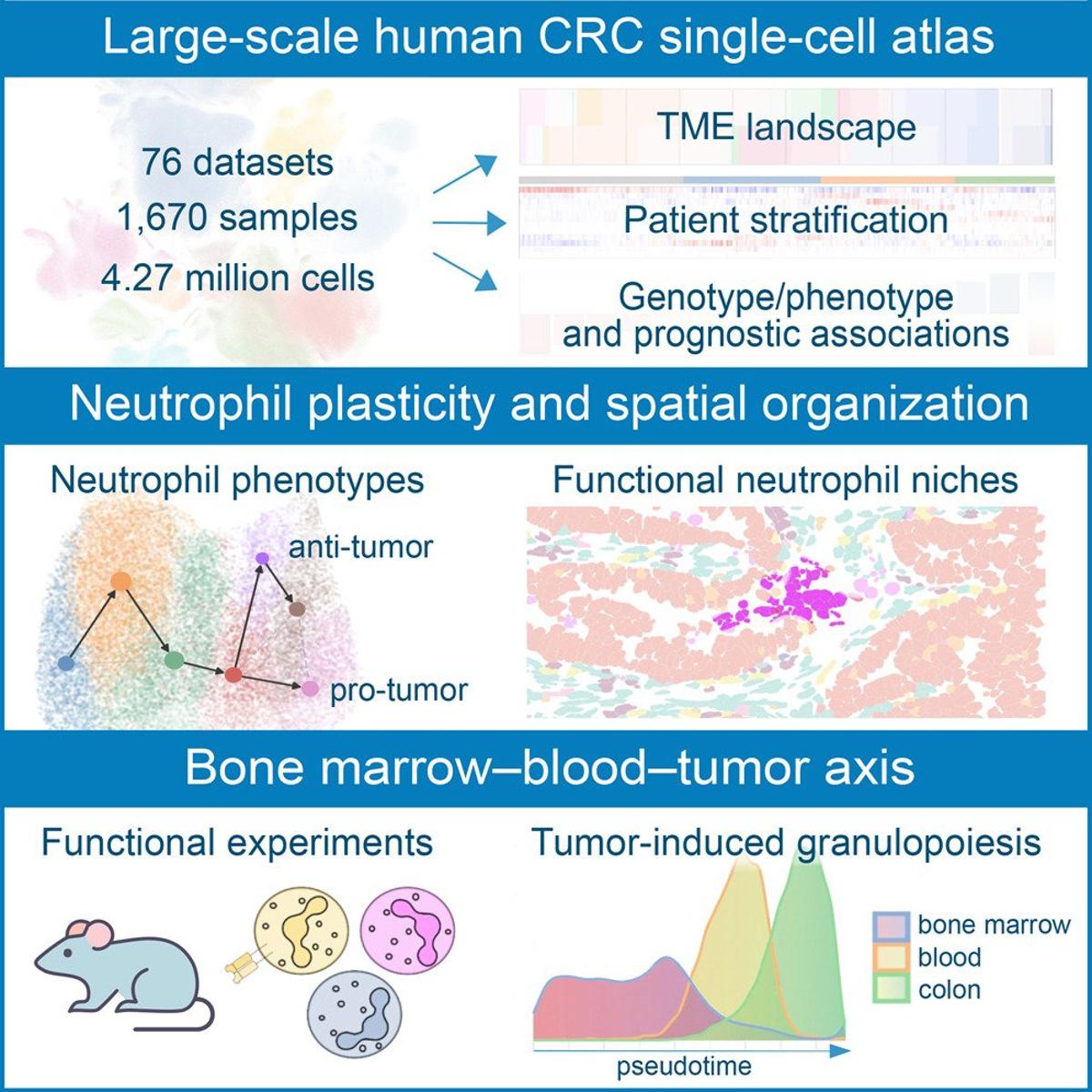
Neutrophil Phenotypes and Spatial Patterns Mapped in Colorectal Cancer
Single-cell integration and multi-modal profiling reveals phenotypes and spatial organization of neutrophils in colorectal cancer https://t.co/0FLusa7vuB https://t.co/8MswWoLhCa
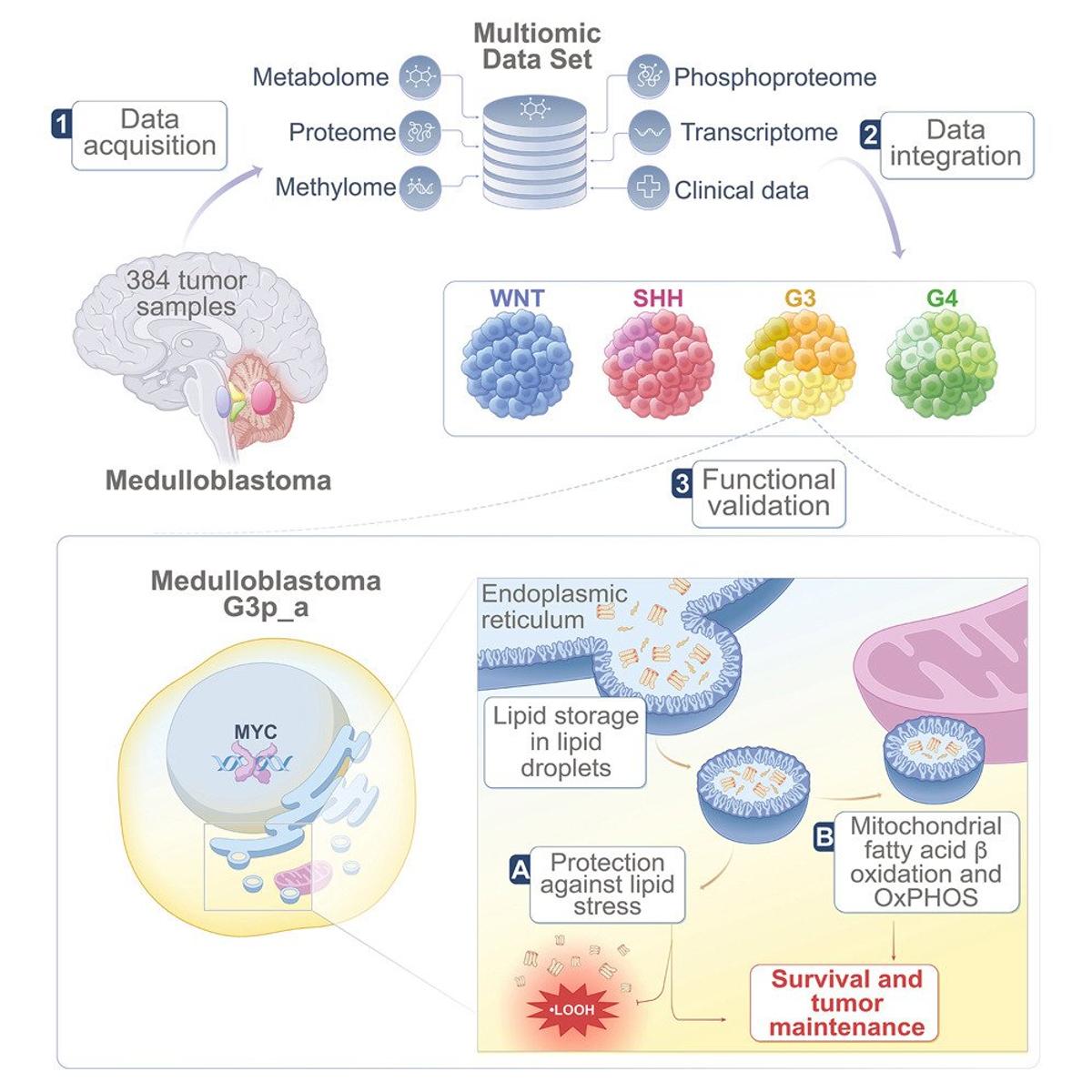
Group 3 Medulloblastoma Shows Diverse Lipid Dependencies
Multiomic integration reveals tumoral heterogeneity of lipid dependence within lethal group 3 medulloblastoma https://t.co/tpjJiRa08t https://t.co/z7aIYerTqO
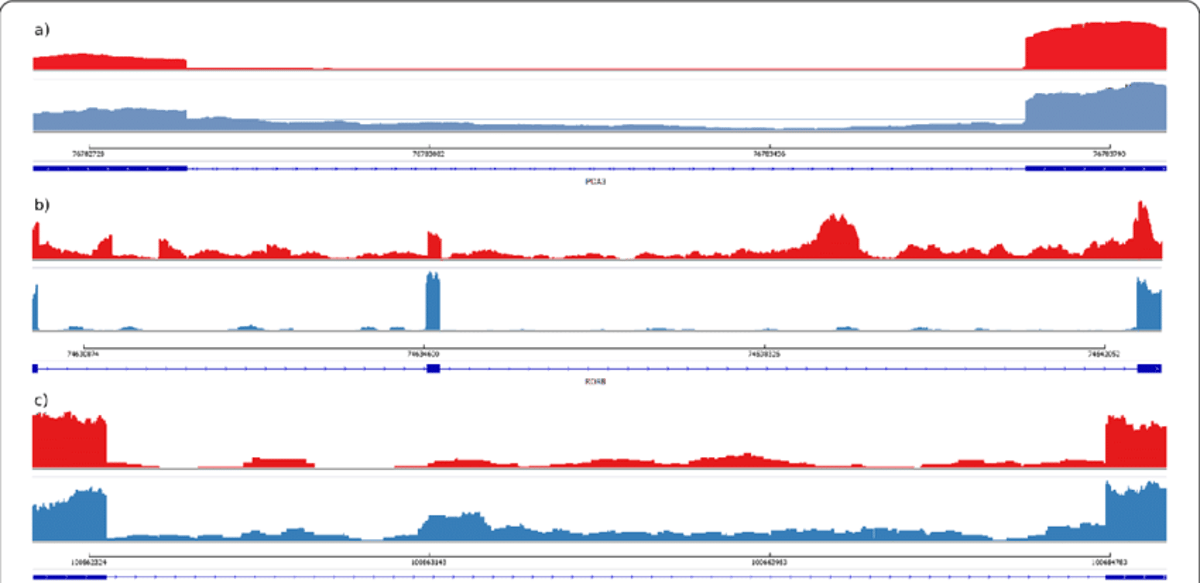
Intronic Reads in Bulk RNA‑seq: Common and Multi‑Faceted
Why are there intronic reads in your bulk RNA-seq data? You're not alone—it's common, and the reasons are more layered than you think. Let’s break it down. 🧵 https://t.co/SNJnohqHUM

V‑plot TF Binding May Be Artifact of Fragmentation
🧵 That V-plot showing transcription factor binding? It might be an artifact. New research shows chromatin fragmentation methods create patterns even on naked DNA. Here's what went wrong. https://t.co/79B5c0ulgD

Claude Code Fabricates Fake ENSEMBL IDs for Gene Plots
1/ Claude Code just hallucinated ENSEMBL IDs for my volcano plot. (it happened last week for real) I asked it to highlight specific genes. The data matrix uses ENSEMBL identifiers. Instead of flagging that it couldn't map the gene symbols, it invented...
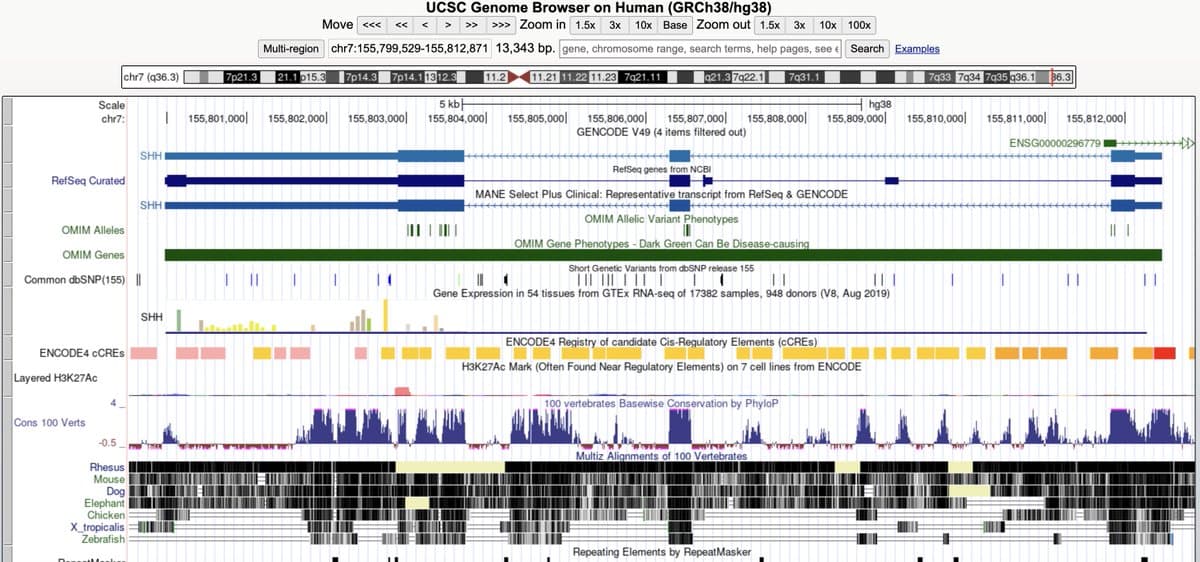
10 Free Bioinformatics Tools to Save Time & Money
🧵 10 free bioinformatics tools you should know in 2026. These will save you time, money, and headaches. https://t.co/t8amdLE4MX
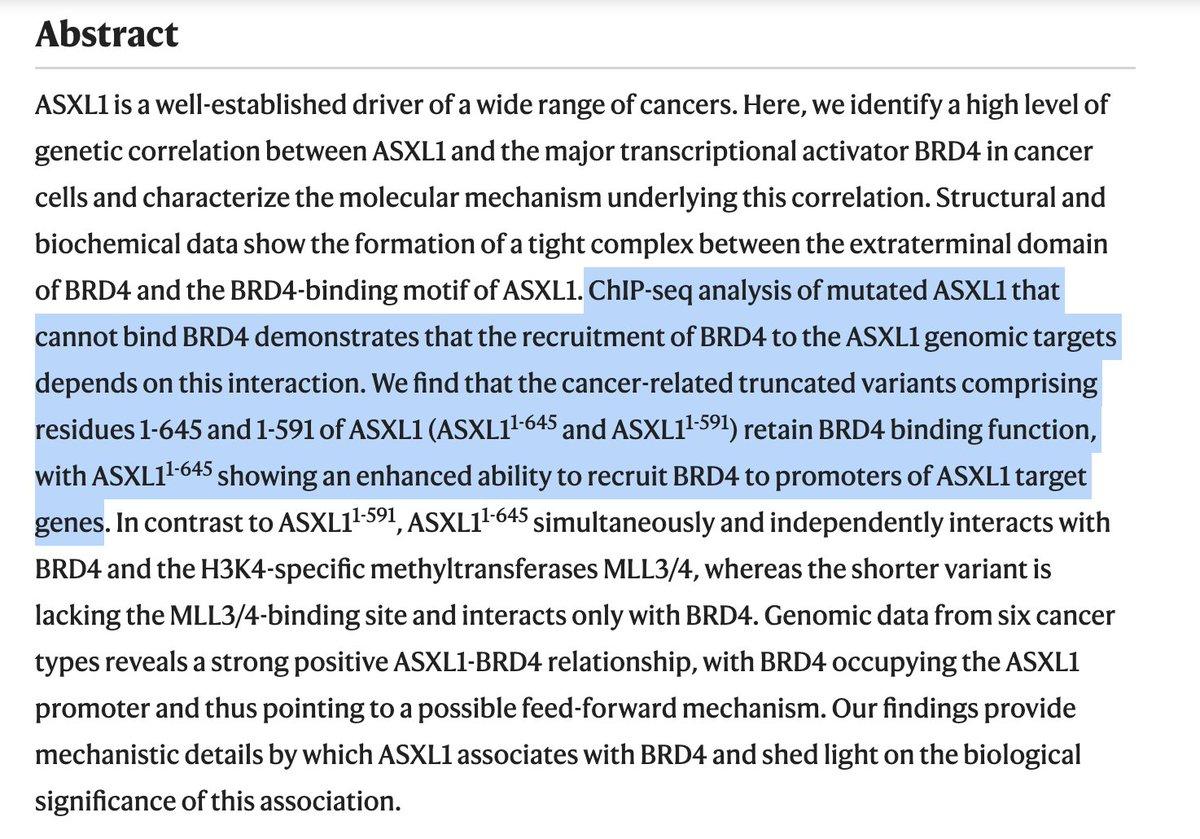

BRD4's Extra‑terminal Domain Drives Recruitment to ASXL1 Targets
Recruitment of BRD4 to the ASXL1 genomic targets depends on the extra-terminal domain of BRD4 https://t.co/sRbcAAMqPx https://t.co/IbD0EM56SS

Retrieve Mouse Gene Lengths with Bioconductor TxDb
🧵 Need gene lengths for every mouse gene? Bioconductor annotation packages make this simple. Here's how to get gene lengths using TxDb.Mmusculus and https://t.co/FiqDAsSPa0.db. https://t.co/VyE36hp3om
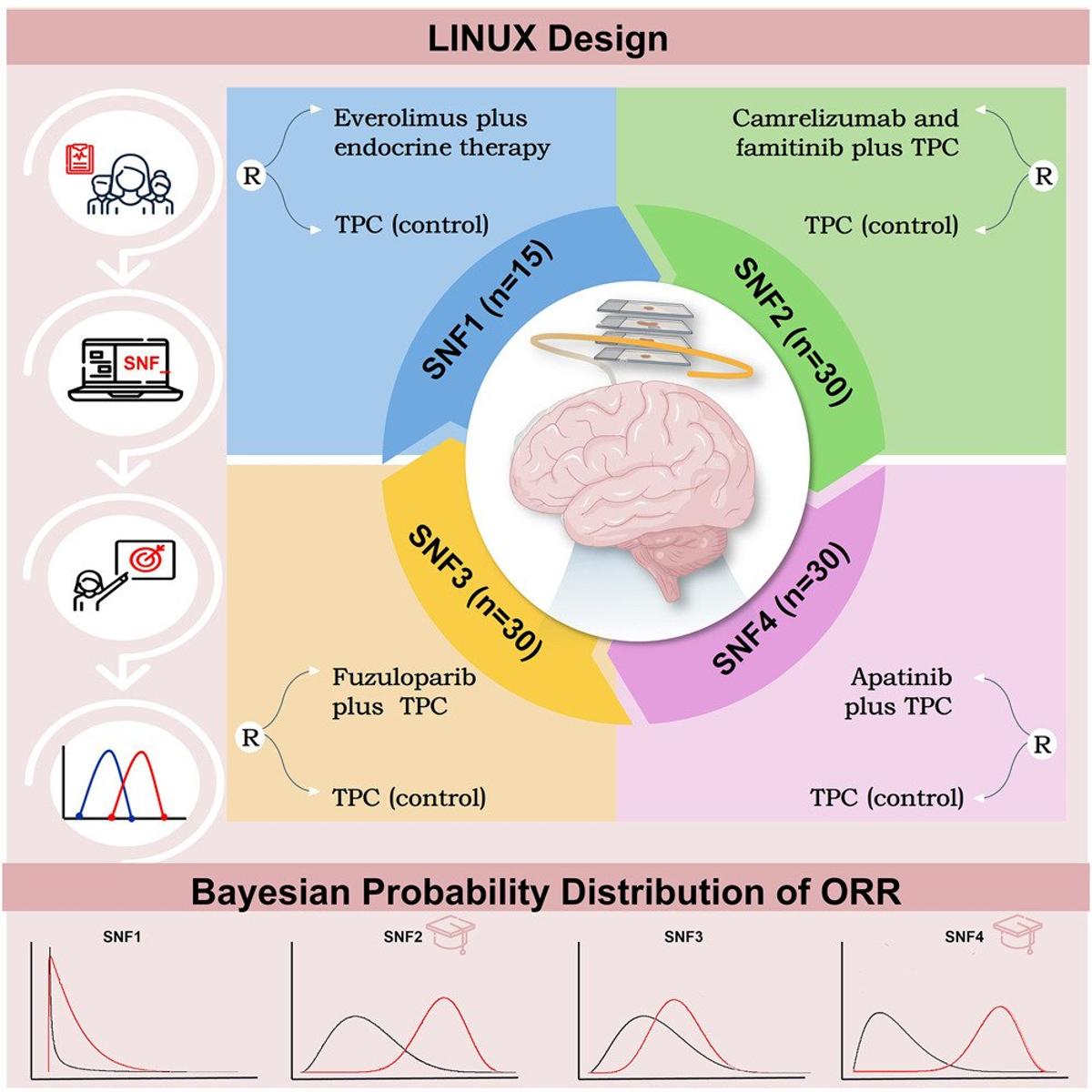
AI Subtyping Boosts HR+/HER2‑ Breast Cancer Therapy
Precision treatment with artificial intelligence assisted subtyping enhances therapeutic efficacy in HR+/HER2− breast cancer: The LINUXtrial https://t.co/UF4Lt3g5QK https://t.co/LWuJYD2qGh

Essential Bioinformatics Curriculum: Top Courses to Master
10 courses for my dream bioinformatics curriculum: 1. Unix Commands with Greg Wilson https://t.co/83VHvTqbMt 2. statistics and R with Rafael Irizarry https://t.co/ofYSIiChoW https://t.co/1GogvjAbAA
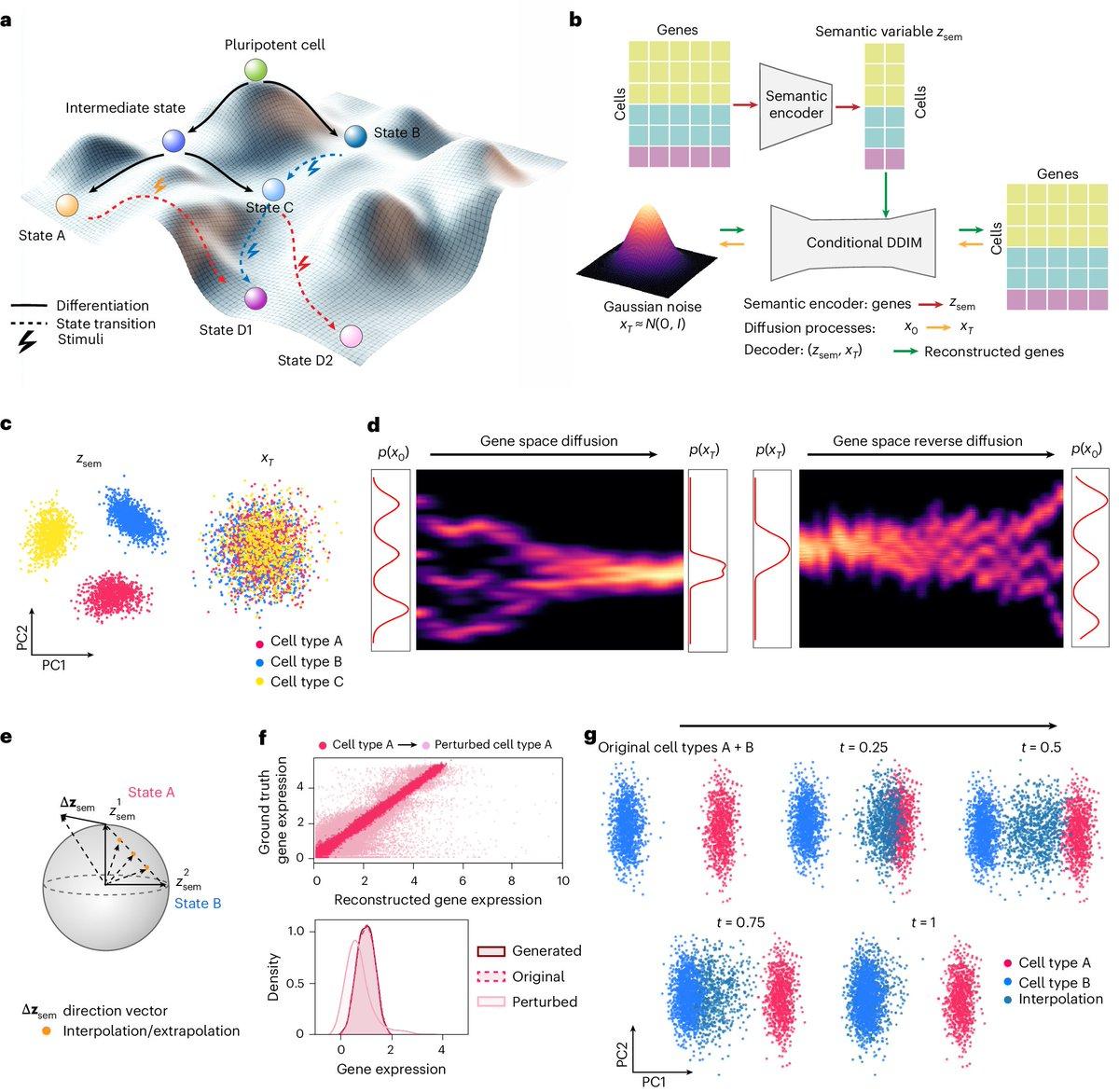
Squidiff Predicts Cell Development and Perturbation Responses
Nature Methods: Squidiff: predicting cellular development and responses to perturbations using a diffusion model from single cell data https://t.co/MqJxhiRJDD https://t.co/cV0IwwFABZ
Pick a Bioinformatics Tool and Just Start
🧵 Stop searching for the "perfect" bioinformatics tool. You're wasting time. Here's why picking something and moving forward beats endless comparison. https://t.co/cffR3dJaoQ

AI Speeds Biology Discovery, Scientists Still Steer Ethics
AI is not taking over biology, at least not now—it's accelerating discovery, not replacing scientists; humans still ask the questions, validate results, and steer ethical choices. https://t.co/l8KCQGNd4Y

Data Hygiene Precedes AI Success in Biotech
1/ You can't bolt AI onto chaos. In biotech, if your data is a mess, your AI won't save you. Build the data strategy first. Here's how. https://t.co/HM7qddrCsC
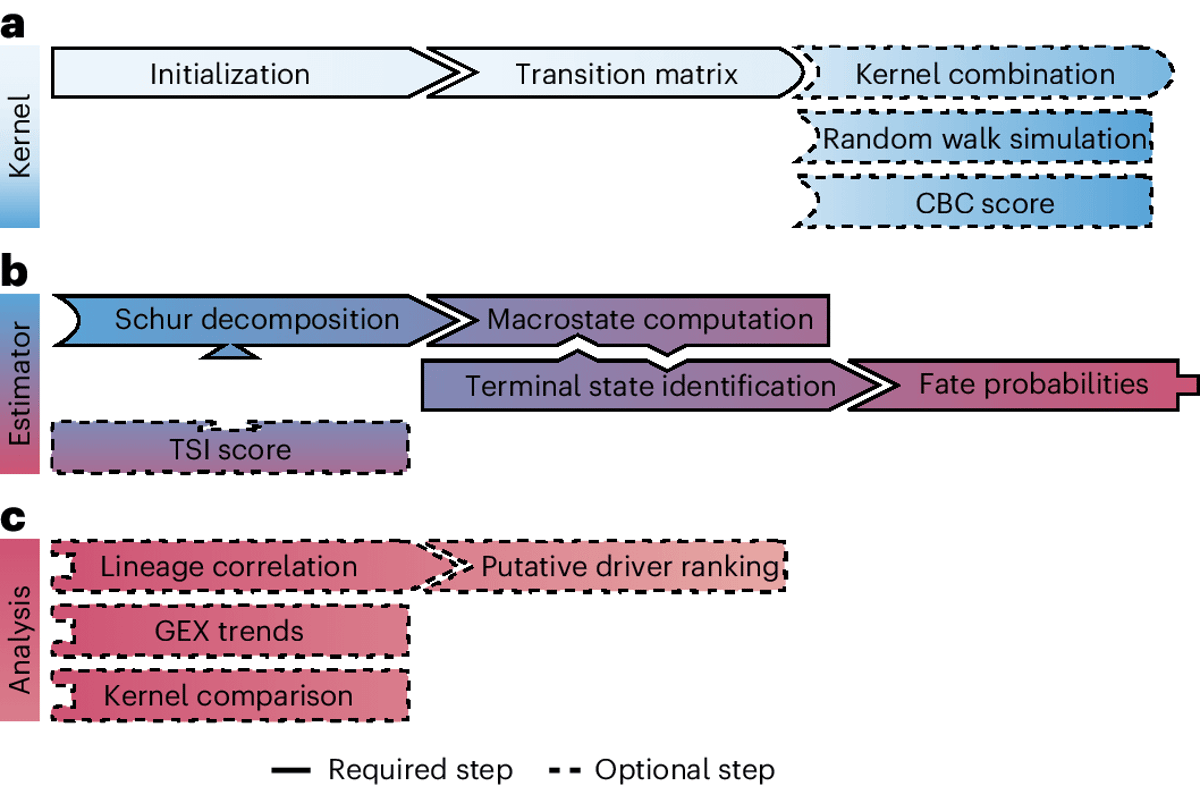
CellRank Maps Trajectories From Destructive Single‑cell RNA‑seq
🧵 Single-cell RNA-seq is destructive. You sequence the cells, they're gone. So how do you reconstruct cellular trajectories? Like tracking stem cells as they differentiate? Enter CellRank. https://t.co/HqHnErHYNy

Stay Grounded Amid Rapid Bioinformatics Advances
🧵Bioinformatics evolves fast. New tech. New data. New analysis. But here's how to stay grounded and not get overwhelmed: https://t.co/ZyxcLRExC6
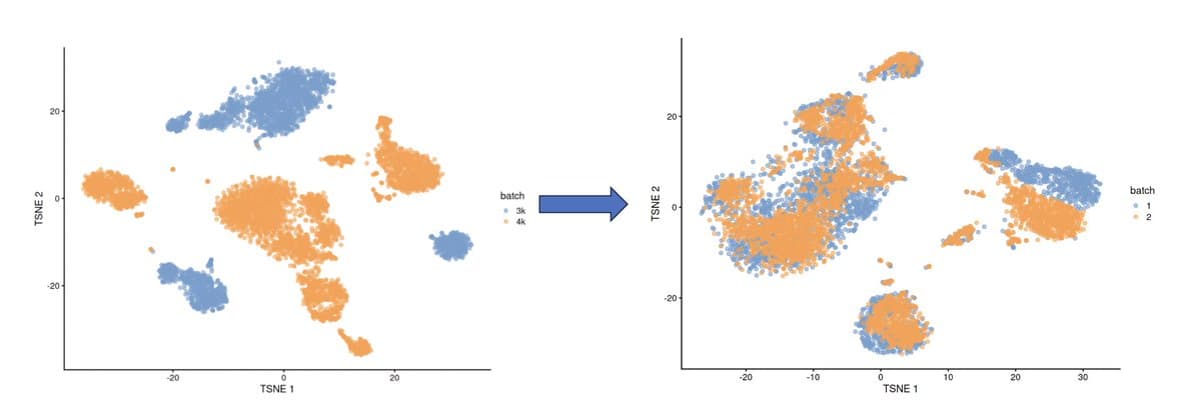
When to Merge Vs. Separate Single-Cell Datasets
🧵 Should you integrate single-cell RNA-seq datasets or not? You've got PBMCs from multiple donors. Merge them—or keep them separate? Let's break it down. https://t.co/s7PmZrvalM