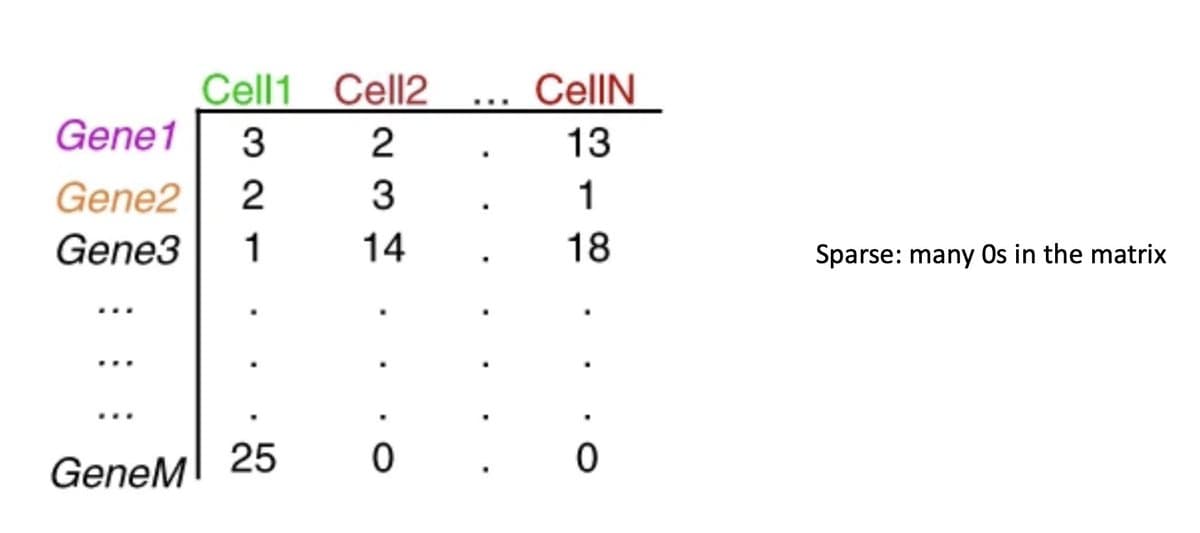
Bioinformatics Demands Matrix Thinking—Most Never Learned It
1/ If you're in bioinformatics, you're staring at matrices all day. RNA-seq? Gene x sample. scRNA-seq? Gene x cell. Everything is a matrix. But I never learned how to think in matrices. And I regret it. https://t.co/ygbY31AfMe
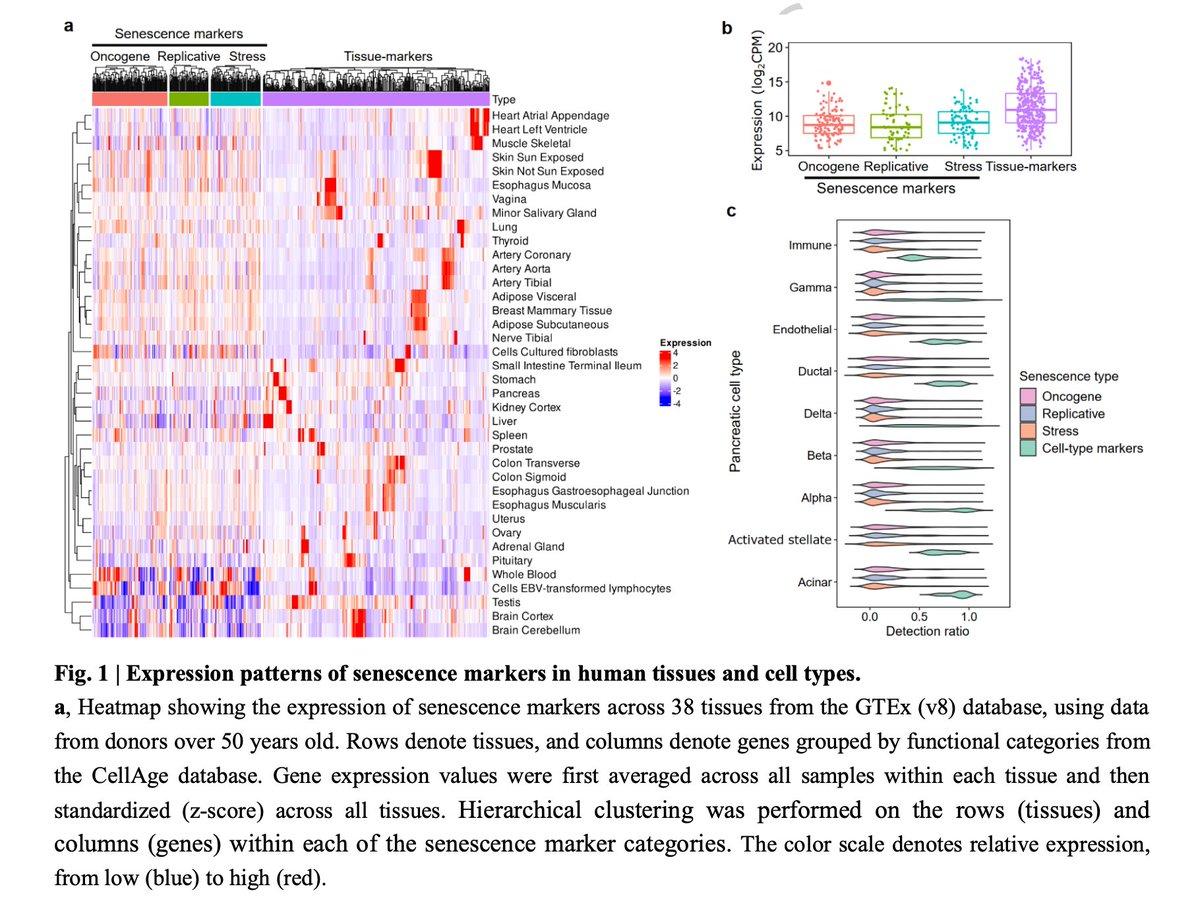
ICE Enables Accurate Senescence Detection From Sparse Single‑Cell Data
ICE: robust detection of cellular senescence from weak single-cell signatures using imputation-based marker refinement https://t.co/PDp6oK5s3W https://t.co/8cPgIQEuID
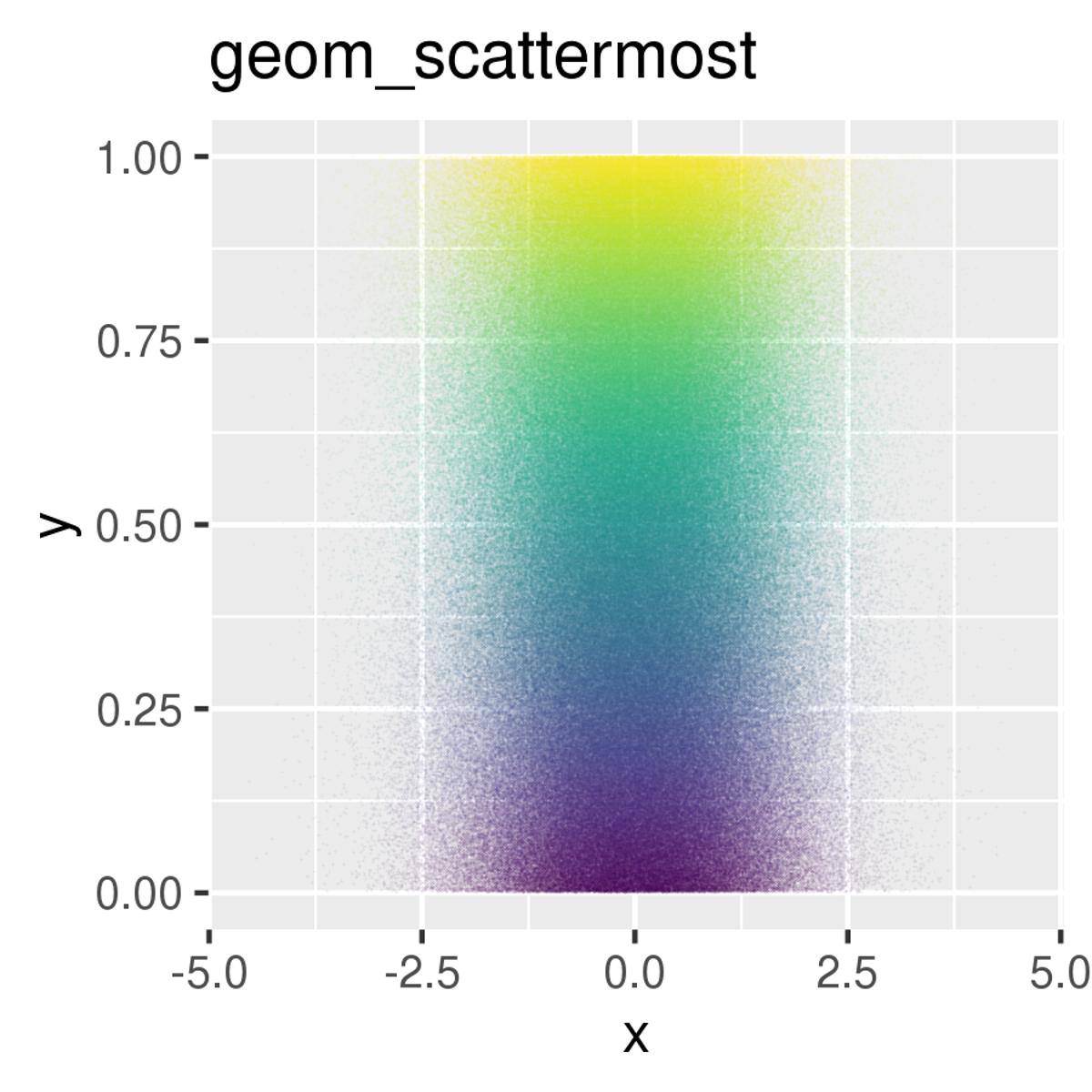
Boost R Scatter Plots with Scattermore for Million Cells
R is slow in plotting tens of thousands of points. How to speed up for a million cell scRNAseq data? check out scattermore https://t.co/tGPHObuK9S https://t.co/TfsRb5V9Xs

Collaboration Between Bioinformaticians and Wet Biologists Unlocks Science
🧵The most underrated superpower in science: Bioinformaticians and wet biologists working together. Here’s why it matters. https://t.co/YNDqsvyu6A

Computational Biology: Beyond Coding to Data Detective Work
🧵 So You Want to Be a Computational Biologist? 1/ Bioinformatics is more than just coding. It’s data analysis, modeling, pipelines, and detective work. Here’s what you must know. 👇 https://t.co/oJxsWkBYzP
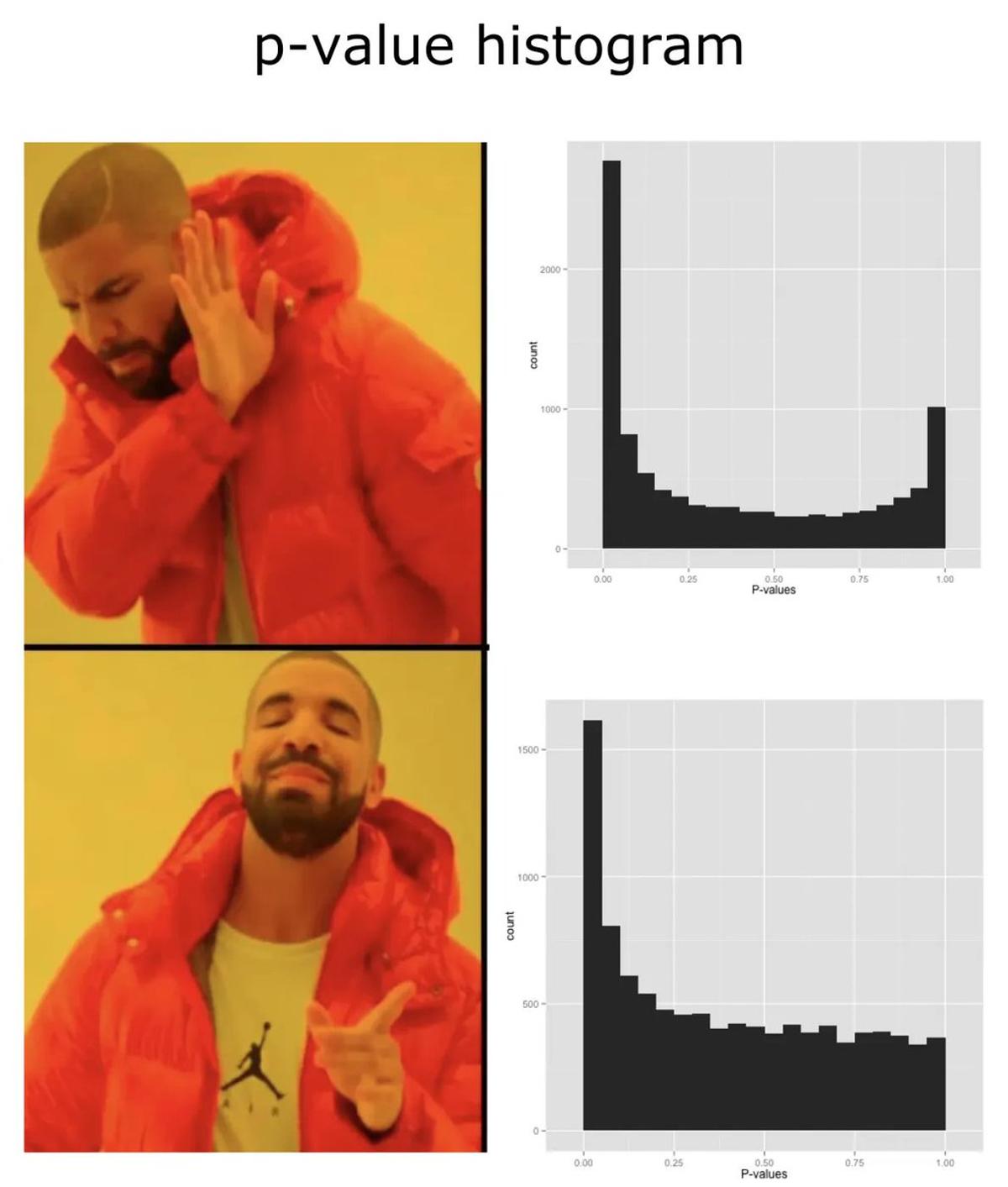
Inspect Your P‑value Histogram After DESeq2 Analysis
How is your p-value histogram look like? chatomics blog post: Downstream of bulk RNAseq: read in salmon output using tximport and then DESeq2 https://t.co/zGjobde9TO https://t.co/aCJaxto5tS
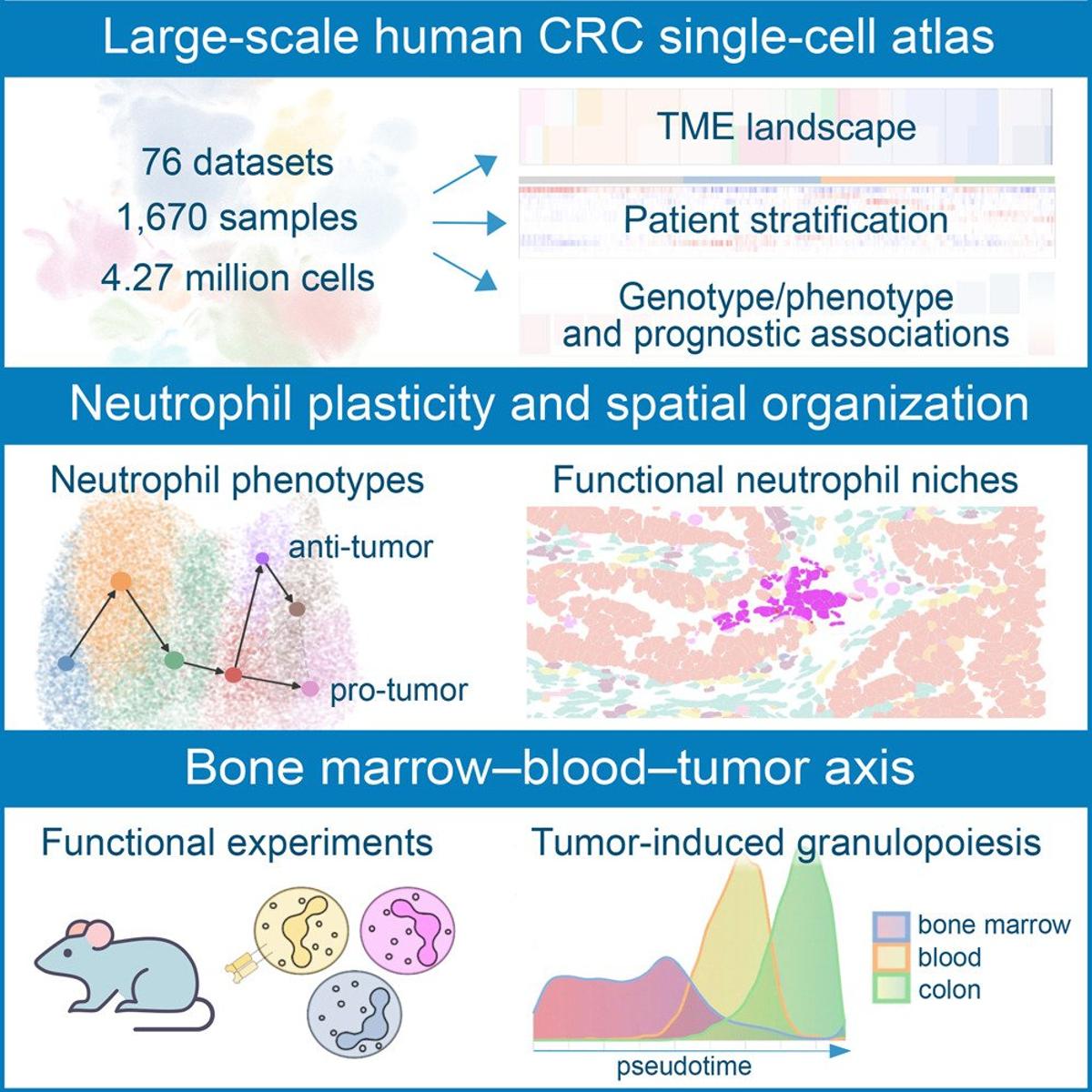
Neutrophil Phenotypes and Spatial Layout Mapped in Colorectal Cancer
Single-cell integration and multi-modal profiling reveals phenotypes and spatial organization of neutrophils in colorectal cancer https://t.co/0FLusa7vuB https://t.co/ETU87KRDXd

AI Tools Like Claude Code Transform Bioinformatics Development
Are bioinformaticians losing their jobs because of AI? After using Claude Code for a couple of months, I'm truly impressed by its ability to write and optimize bioinformatics tools. https://t.co/ojh5bv09fI

Bioinformatics Mastery Stalls Due to Undocumented Tacit Knowledge
1/ Bioinformatics takes years to master. Not because it’s hard. But because so much of what matters… no one writes down. Let me explain https://t.co/EEboWOxB5C

Solid Foundations Beat AI Hype in Bioinformatics
1/ AI won’t save sloppy science. Before you dive into deep learning, master your foundations. Here’s why basic bioinformatics still rules 🧵 https://t.co/nPbaEK3hGf

MAPK‑RUNX1 Axis Controls SMARCA
Spatiotemporal control of SMARCA5 by a MAPK–RUNX1 axis distinguishes mutant KRAS-driven pancreatic malignancy from tissue regeneration https://t.co/eshLHVpD8I https://t.co/mZj4F2qeGp

Curiosity and Mistakes, Not Brilliance, Build Expertise
After 13 years of analyzing sequencing data, I am an "expert" in doing it. I gained those experiences not because I am smarter, but simply because I am curious I made more mistakes, and encountered more problems.

CRISPR/dCas9 Epigenome Editors Show Complex On‑off Target Relationship
Comprehensive profiling of CRISPR/dCas9 epigenome editors indicates a complex link between on and off target effects https://t.co/sEH5GH08Hh

Suspect Mycoplasma Contamination When Human Mapping Fails
Your cell line sequencing data isn’t mapping well? Before blaming the aligner—check if you're sequencing bacteria instead of human. Let’s talk about mycoplasma contamination. https://t.co/y8Ydk0i248
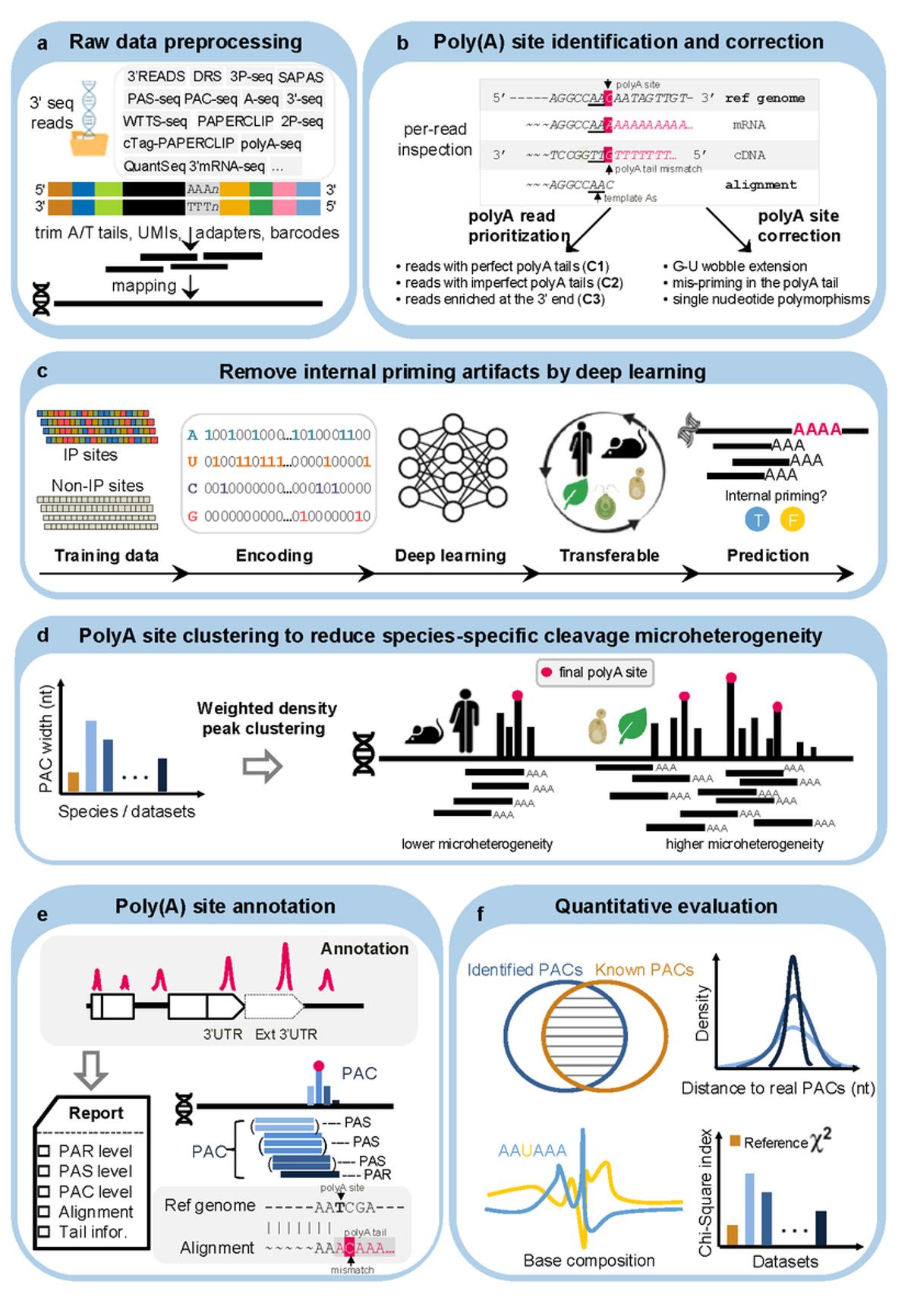
PolyAseqTrap Enables Universal Genome-Wide polyA Site Mapping
PolyAseqTrap: a universal tool for genome-wide identification and quantification of polyadenylation sites from different 3′ end sequencing data https://t.co/pwEf2GeYjb github https://t.co/gpfYhyMoX3 https://t.co/oyA5MBFutE

Master NGS Pre‑Processing: Raw Data Understanding Is Crucial
1/ Many bioinformatics students don’t know much about NGS pre-processing. But trust me, understanding the raw data is essential. Here’s why. https://t.co/WjT1I9haD8

Turn RNA‑seq Dimensionality Chaos Into Clarity
1/Ever feel buried under 20,000 genes and no clue where to start? That’s the curse of dimensionality in RNA‑seq. Let’s turn chaos into clarity. https://t.co/0U3wdzUsMm

Bioinformatics Analysis Time Varies; Depends on Many Factors
“How long will the bioinformatics analysis take?” If you’ve ever been asked that… you know the answer: It depends. Here’s why: 🧵 https://t.co/PG7Xaj3oxf

CHIP in Blood Sequencing Skews Mutation Detection
1/12 What is “CHIP” in blood sequencing, and why does it mess up mutation calls? Not every mutation is created equal. Quick explainer for oncologists, cardiologists, and genomics folks 👇 https://t.co/nBEWTsdlz3

Academic Software's Hidden Risk: Single‑grad Maintainer Dependency
Academic software is a quiet risk in many analysis pipelines. A lot of core tools are maintained by one grad student whose incentives don't include long-term upkeep. https://t.co/ec4R4kDi84

Bioinformatics Needs Nuance, Not Hard Cutoffs
You can not do Bioinformatics with hard cutoffs and thresholds Let me tell you something real: Biology is more about p-values. It’s messier than that. 🧵 https://t.co/iewqWMvU6D

Understanding Login vs Interactive Shells in Bioinformatics
Bioinformatics is more about biology and code. It's shells and sessions, exports and environments. Ever been confused by “login shell” vs “interactive shell”? This thread is for you. 🧵 https://t.co/DfB8BEztiD

Model Failures Stem From Data, Not Algorithm Power
1/ You think your ML model fails because it’s “not powerful enough”? No. It’s your data. Garbage in, garbage out. Here’s what most AI scientists miss when using public RNA-seq or single-cell data 👇 https://t.co/dVezFRLHkW
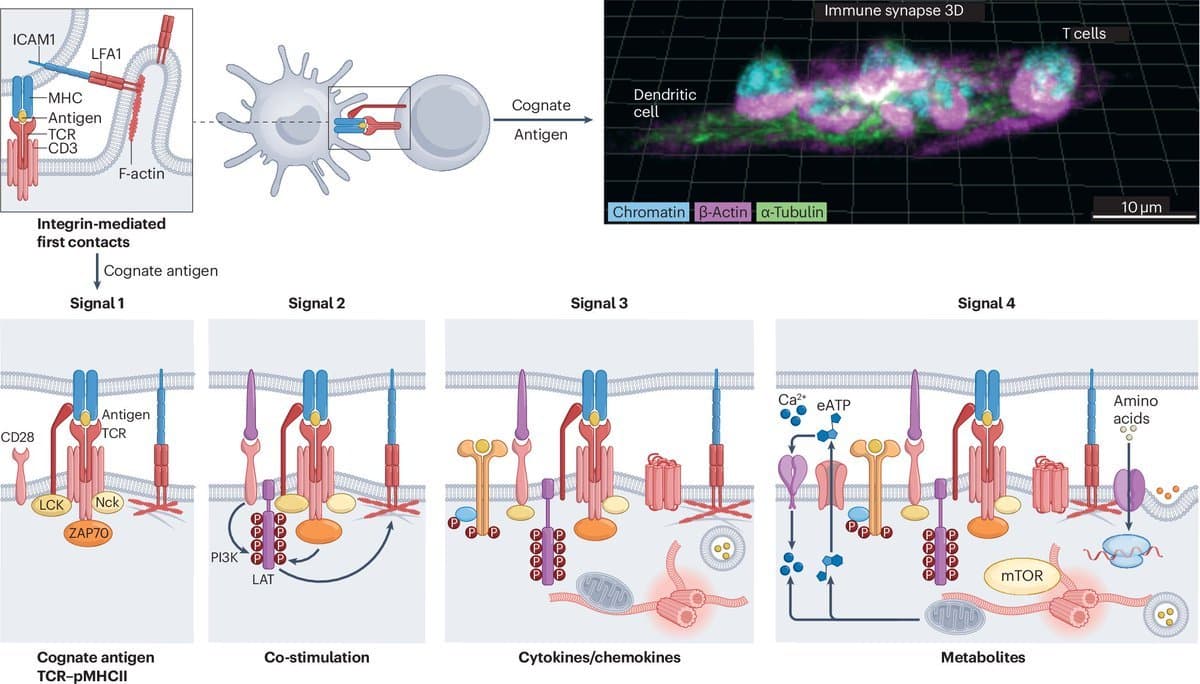
Immune Synapse Crosstalk Directs T Cell and Dendritic Cell Functions
How crosstalk at the immune synapse shapes T cell and dendritic cell biology https://t.co/DtqWi1PWhU https://t.co/Yw3sO2UgSX

Inherited Messy Bioinformatics Code? Learn Better Practices
You inherit someone else’s bioinformatics code. No comments. No structure. Variable names like x1, foo, temp2. And now it’s your problem. Let’s talk about that experience—and how to do better. https://t.co/RqCDNPwhMD
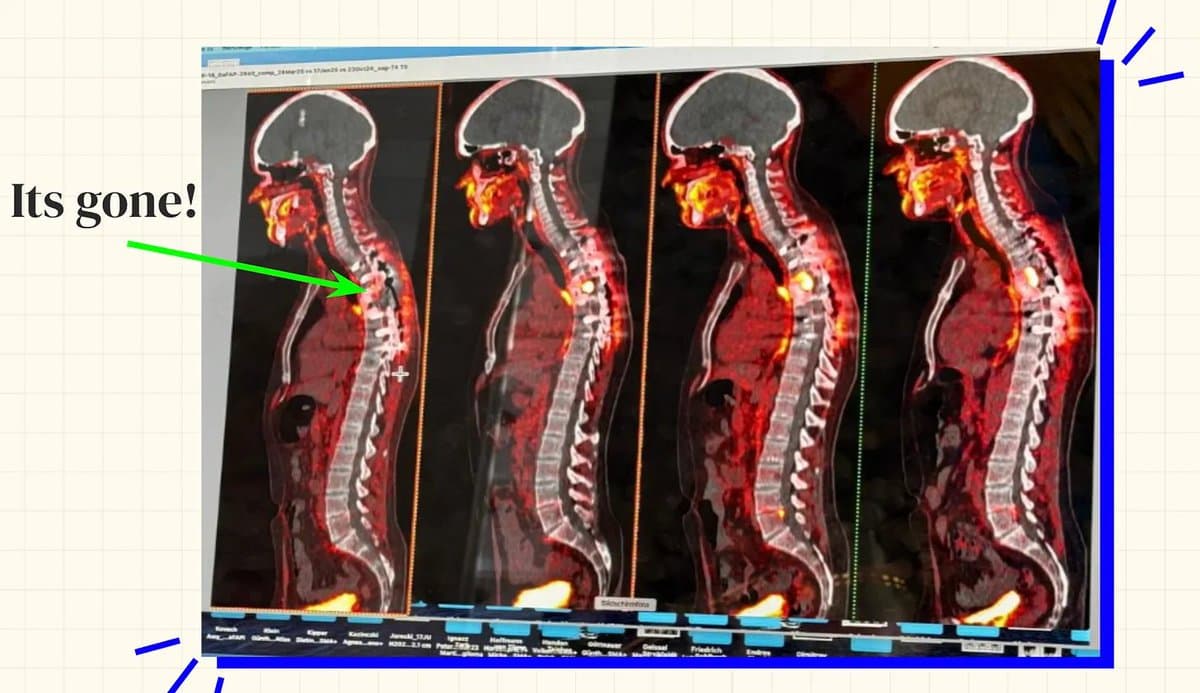
Billionaire Sid's Cancer Escape Highlights Access Inequality
Fantastic read. How a billionaire saved himself (for now) from cancer. Not everyone has resources like Sid. https://t.co/73Oj3KF4RP This imposes a bigger question on how we can bring therapeutics to every patient. https://t.co/NyekRQUoND

Batch Effects Create False Cross‑Chromosome Variant Signals
Batch Effects Are Hiding in Your Variant Calls I thought my QC was solid. Then I found thousands of "variants" that weren't real. The signal? Variants on different chromosomes showing linkage disequilibrium. That's impossible in real biology. https://t.co/QK4BVSpbsh

TSniffer Enables Unbiased De Novo RNA Editing Detection
TSniffer: unbiased de novo identification of RNA editing sites and quantification of editing activity in RNA-seq data https://t.co/uQRXdipF88 https://t.co/yeOxSvxrl2

Learning Skills Turns AI Advice Into Useful RNA‑seq Tools
Claude keeps suggesting outdated tools for my RNA-seq analysis. Then I learned about skills. Now it actually helps instead of creating work. https://t.co/yiFbouglo2

Sequencing Artifacts Masquerade as Rare Disease Mutations
1/ Your genome report says you have a disease-causing mutation. Reanalysis 13 months later: it was a sequencing artifact. MedSeq found 164 "rare variants" appeared in >10% of their patients. Population databases missed them all. https://t.co/S0hc1TRr18

KNN in scRNA‑seq: More Art than Science
Everyone talks about KNN (K nearest Neighbor) like it’s a simple algorithm. But in practice, especially in single-cell RNA-seq—it’s art, not science. https://t.co/BCreDmtKwZ

Highly Cited Cancer AI Models Rarely Reach Patients
1/ Your cancer prediction model has 1,000 citations, but It's never been used on a patient. "Some models are wrong, yours are useless." A Cambridge researcher analyzed why most clinical AI tools die in academic journals instead of helping people. https://t.co/mIXS32qDr3
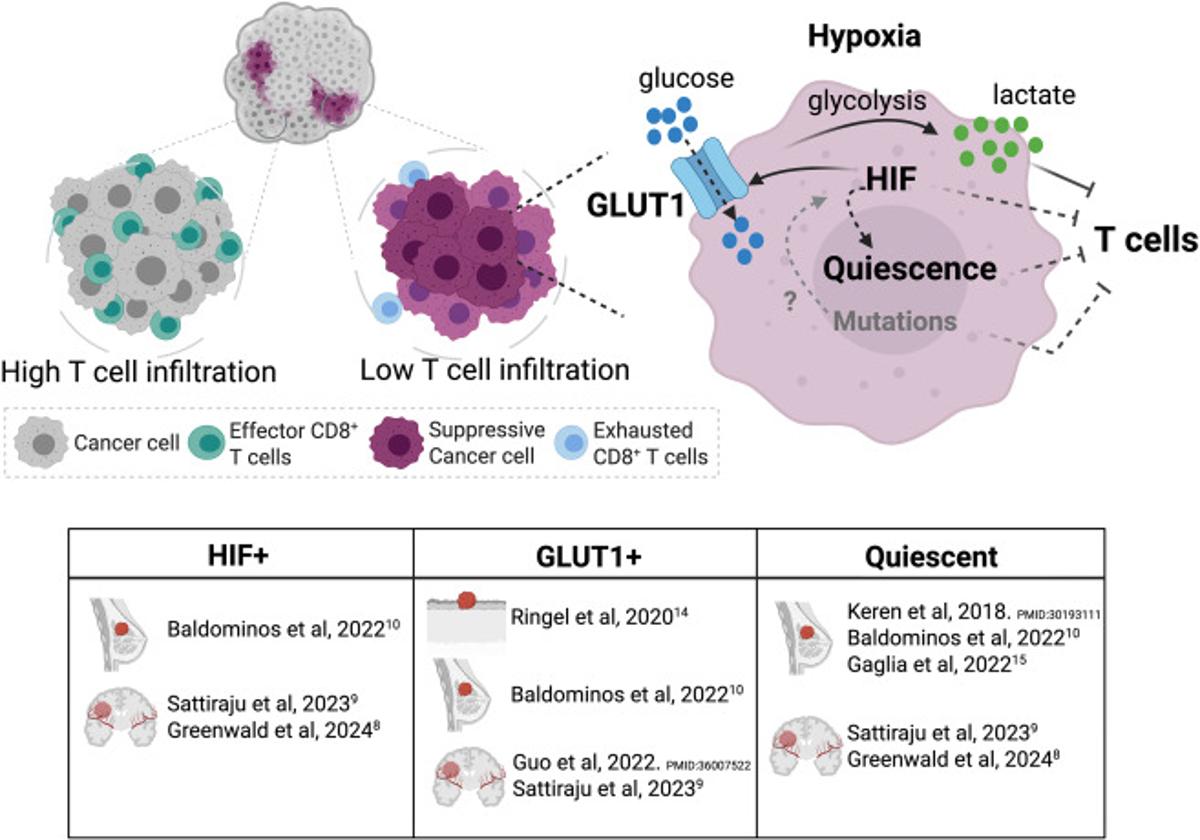
Mapping Tumor‑Immune Spatial Interactions in Solid Cancers
Decoding the spatial dynamics of tumor and immune cell interactions in solid cancers https://t.co/AHgt77Ee2t https://t.co/1Xxkt45JlD

Use Repetition to Slash Bioinformatics Analysis Time
Are you ready to level up your bioinformatics skills? Let’s talk about repetition—a key concept that can save you hours in real-world data analysis. https://t.co/2Dtc9hz46e
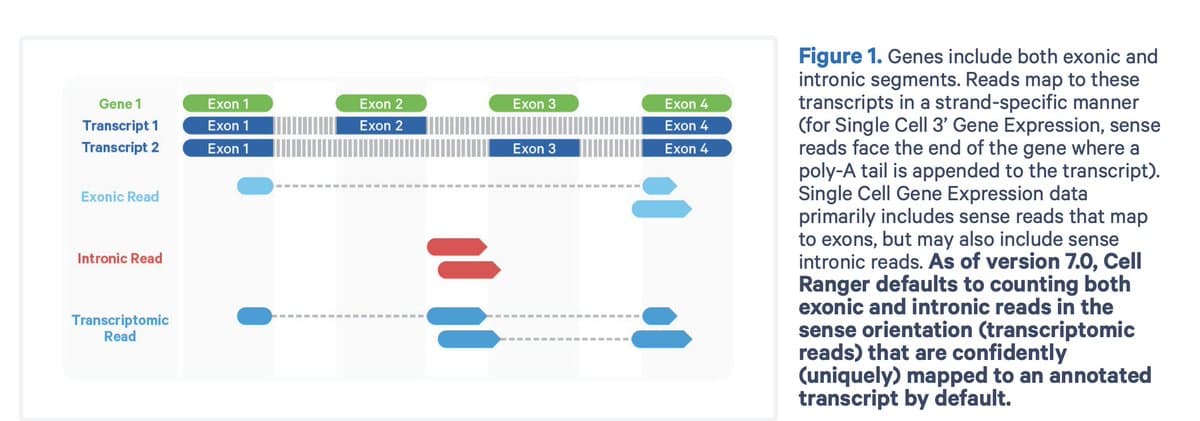
Intronic Reads in 10x 3′ UMI Data Explained
You’re analyzing 10x Genomics single-cell RNA-seq and notice lots of intronic reads. Wait—wasn’t this a 3′ UMI-based assay for mature mRNA? Let’s unpack why introns show up—and why they matter. 🧵 https://t.co/cDeb8dfLAS
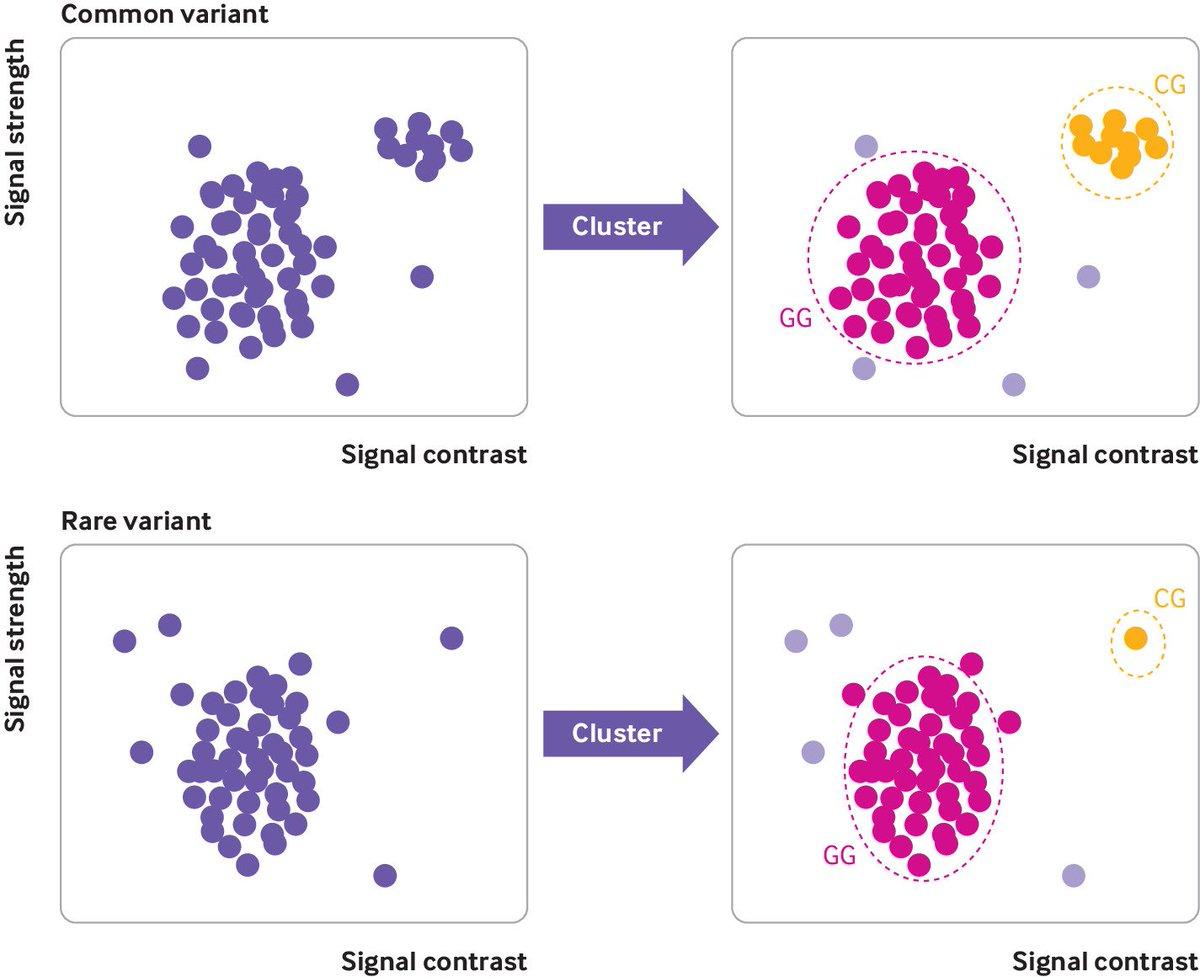
Study Finds Frequent BRCA1 Test Errors, Risking Unnecessary Surgeries
1/ Women scheduled surgery after being told they had rare BRCA1 variants. The genetic test was wrong. University of Exeter analyzed 50,000 samples to find out how often this happens. The results should worry anyone who's downloaded their 23andMe raw data. https://t.co/F1QALvi4H4

Stop Wasting Hours Matching Sample IDs Across Assays
1/ How many hours do bioinformaticians lose matching sample IDs across assays? Too many. And it’s avoidable. Let’s talk about why this happens—and how to stop it. https://t.co/ZIcF1bUzFF

DUD‑E Bias Inflates Deep‑learning Virtual Screening Results
Hidden bias in the DUD-E dataset leads to misleading performance of deep learning in structure-based virtual screening https://t.co/NEAe3zdQrq
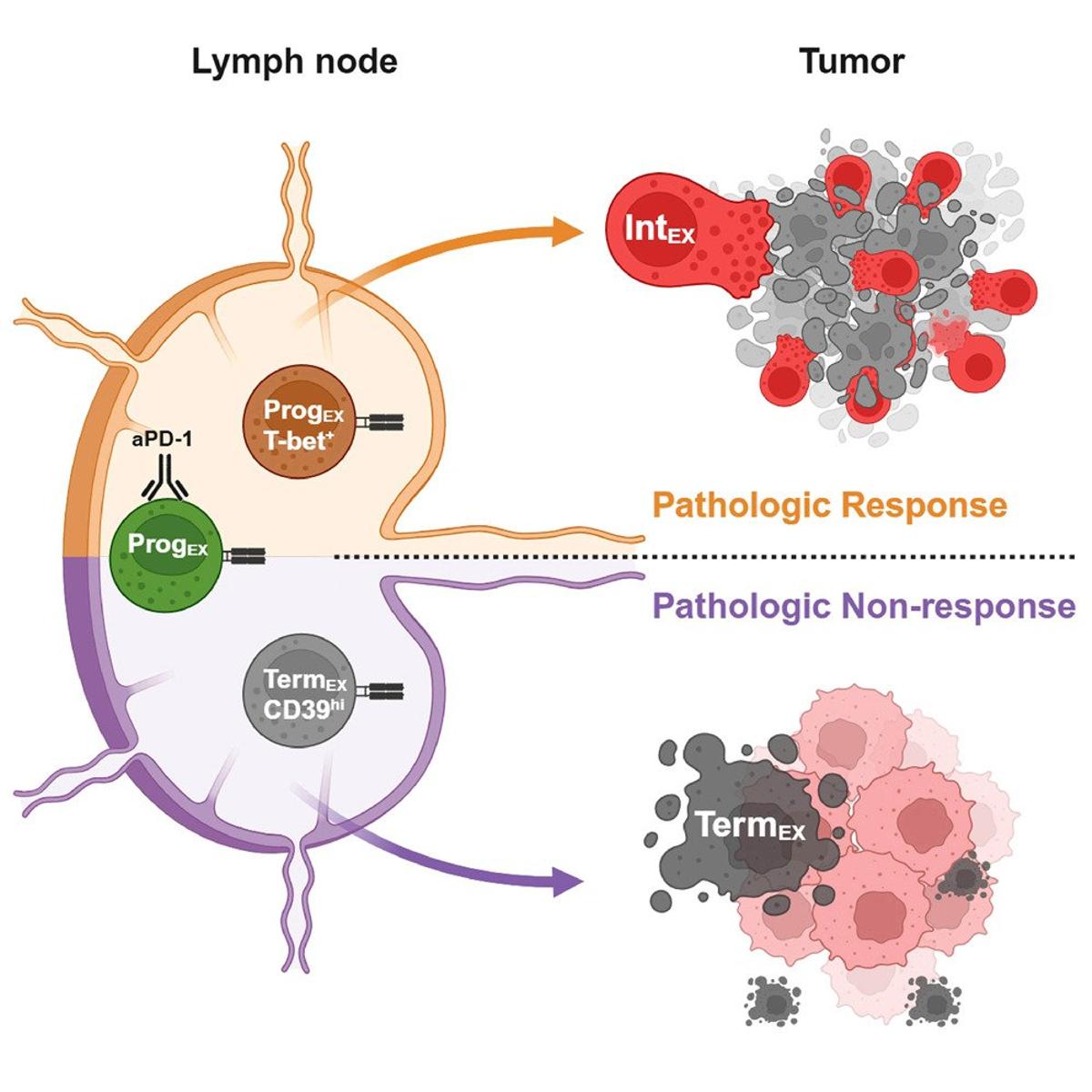
T‑bet+ CD8 T Cells Predict Neoadjuvant PD‑1 Response
Antigen-specific profiling identifies T-bet+ melanoma-specific CD8+ T cells associated with response to neoadjuvant PD-1 blockade https://t.co/VSwMpdqjqO https://t.co/DXc8P28oIZ

One Year In, Code Works—Beware Self‑Deception
1/ A year into bioinformatics, your code starts to work. But that’s also when it gets dangerous. Because now you can fool yourself. https://t.co/7QCMSpI30A

SIAE Variant Linked to Autoimmunity Lacks Control Evidence
1/ Eight patients had a genetic variant. Zero controls did. That 2010 Nature paper claimed SIAE variants increased autoimmune risk 8-fold. The mouse data was clean. The functional assays checked out. https://t.co/tFKXjXEr1y

Longevity Gene Claim Debunked: Study Proves Wrong
1/ A 2010 Science paper claimed they'd found genetic variants strongly tied to living past 100. New York Times covered it. Media went wild. One problem: The results were wrong.
Bioinformatics Blends Code with Intuition for Insight
1/ Bioinformatics isn't just code. Intuition plays an important role too. You run the stats, but you feel when something’s wrong. That feeling is a clue. https://t.co/pV5SxvYuFL
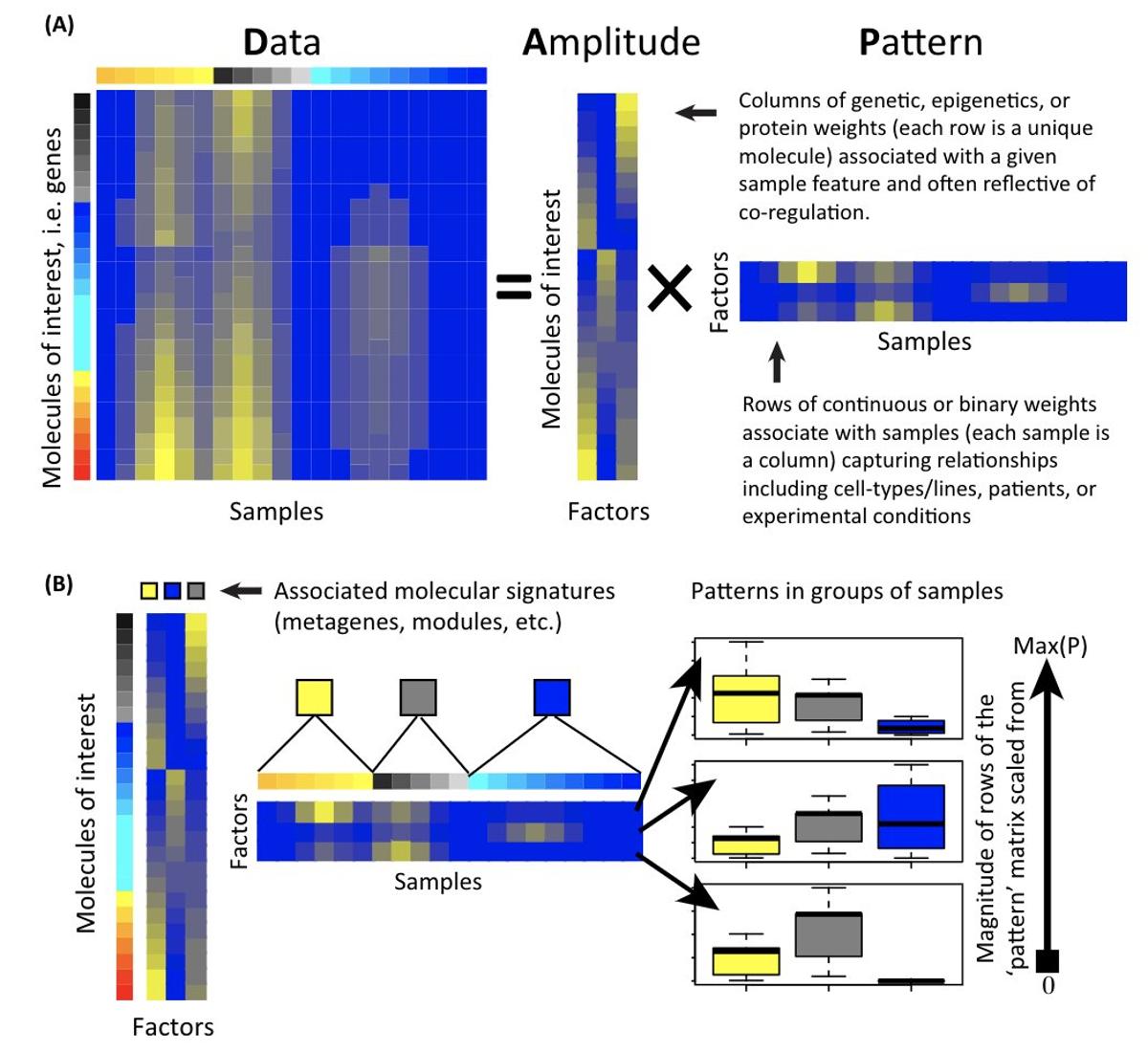
Apply NMF to Single‑Cell RNA‑seq with Our Tutorial
Non-negative matrix factorization is a commonly used technique in genomics data analysis. Read my tutorial on how you can use it for single-cell RNAseq data https://t.co/2SA1JdLfkT https://t.co/6kI1gcqyOe
Twitter Transformed My Bioinformatics Career and Keeps Me Current
Bioinformatics is a fast-moving field, how to stay current? 👇 The answers are different in different times. I read Stephen's post around 2012 and I hopped on Twitter; followed a bunch of bioinformaticians, Journals and professors. It changed my career trajectory. https://t.co/QYV5vKFQgP
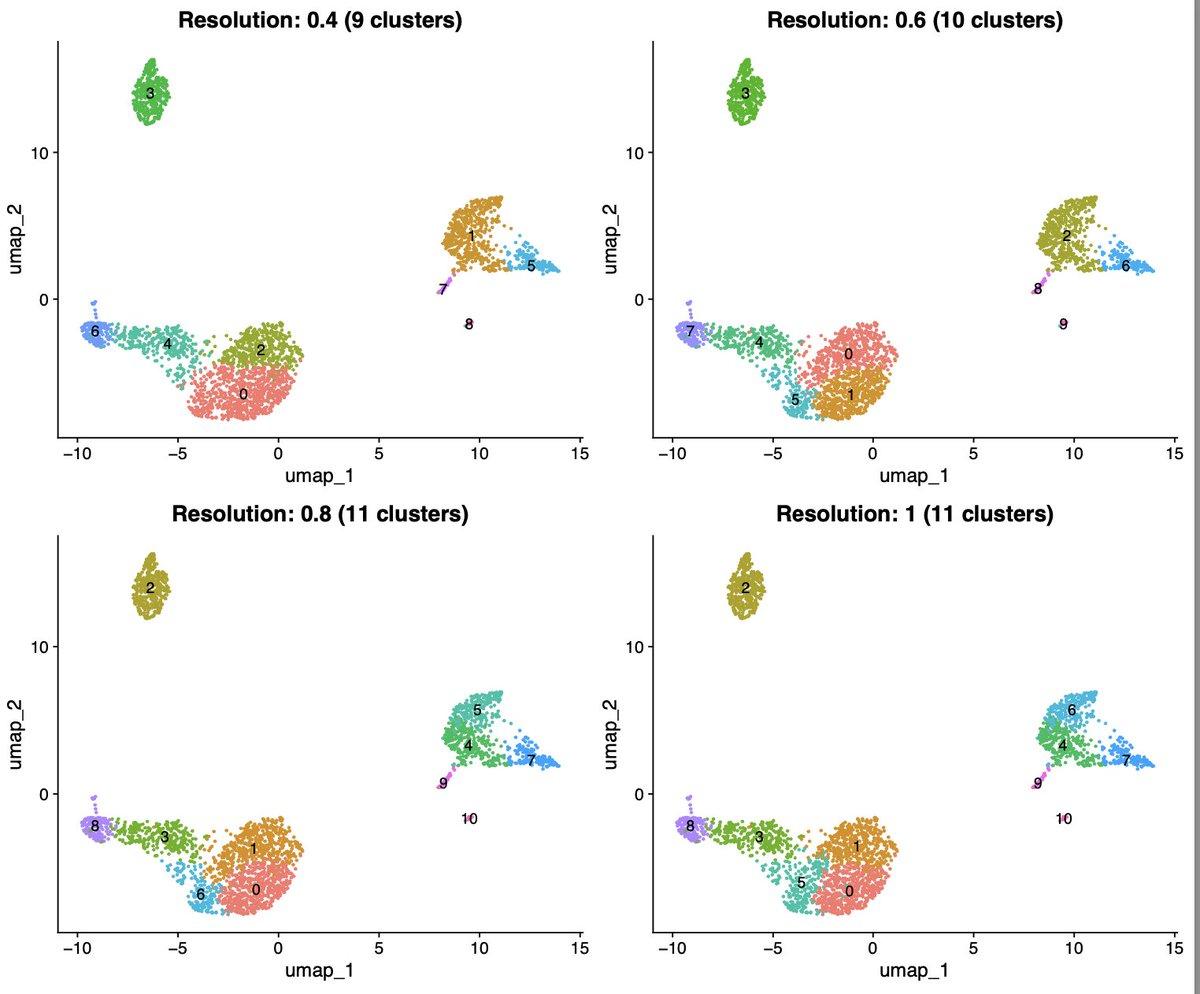
AI Explains Single‑cell Clustering, Not Just Runs It
1/ I've analyzed dozens of single-cell datasets. I still google "Seurat clustering parameters" every single time. Last week I tried something different. I asked Claude Code to explain my clustering results instead of just generating them. https://t.co/AMQpheqhYC
Using Claude Code Wrong? Real Genomics Workflow Works
1/ Bioinformaticians get mediocre results with Claude Code and blame the tool. You are doing it wrong. Here's what actually works for genomics analysis: https://t.co/GDOq7SPzla

Six Essential Bioinformatics Workflow Tools Simplify Analysis
6 links on workflow to make your life easier 🧵 Bioinformatics analysis involves a lot of steps, 6 links on workflow to make your life easier: 1. over hundreds of workflow tools and engines https://t.co/R29TTEYSMB

AI Rebuilt My RNA‑seq Pipeline in Minutes
1/ I wasted hours debugging an RNA-seq pipeline. The next day, I rebuilt it in 45 minutes using Claude Code.
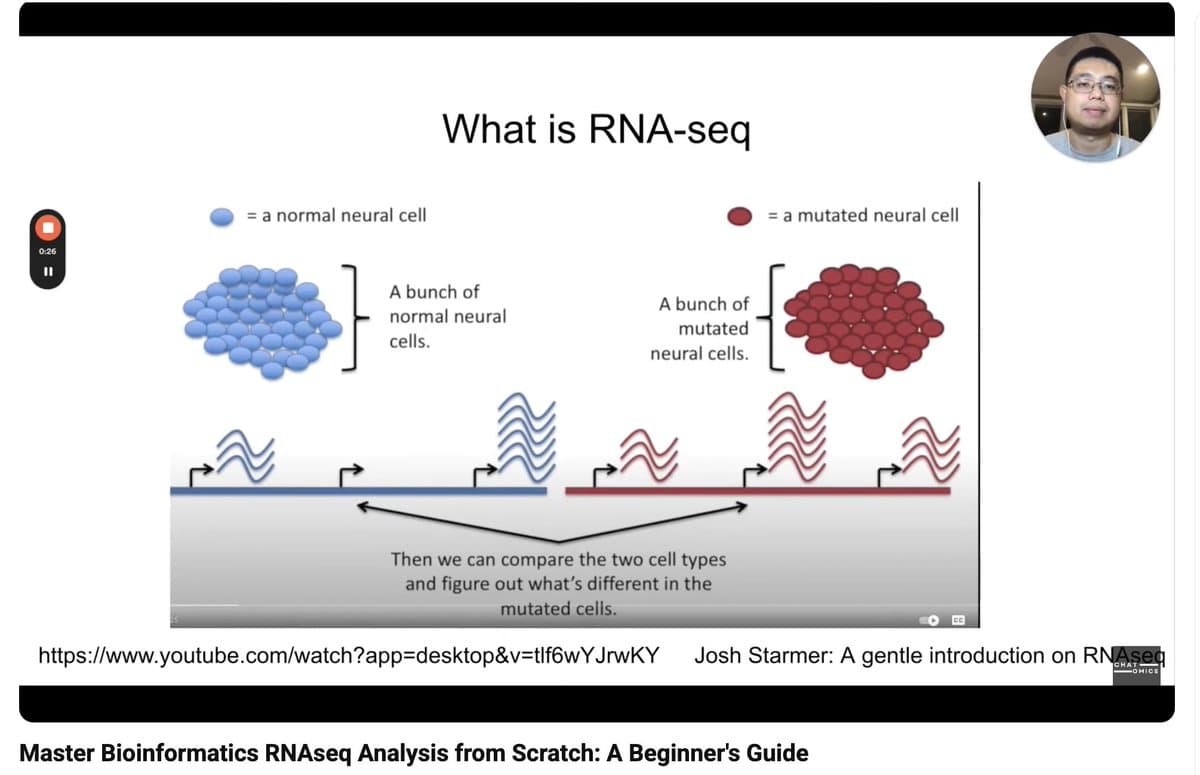
Free Tutorial: From FASTQ to GSEA in Bulk RNA‑seq
Free tutorial: bulk RNAseq analysis from fastq to GSEA analysis (watch the full playlist) https://t.co/v0UHRSqJ63 https://t.co/LtdxPNYG5Y